Analysis of diagnostic tests without a gold standard reference using multiple statistical approaches. Implements Latent Class Analysis (Hui & Walter, 1980), Bayesian methods (Joseph et al., 1995), and composite reference standards for estimating test performance when no perfect reference test exists.
Usage
nogoldstandard(
data,
clinicalPreset = "none",
test1,
test1Positive,
test2,
test2Positive,
test3 = NULL,
test3Positive,
test4 = NULL,
test4Positive,
test5 = NULL,
test5Positive,
method = "all_positive",
bootstrap = FALSE,
nboot = 1000,
alpha = 0.05,
verbose = FALSE
)Arguments
- data
The data as a data frame.
- clinicalPreset
Predefined clinical scenarios with optimized settings and method recommendations.
- test1
First diagnostic test variable.
- test1Positive
The positive level for Test 1.
- test2
Second diagnostic test variable.
- test2Positive
The positive level for Test 2.
- test3
Third diagnostic test variable (optional).
- test3Positive
The positive level for Test 3.
- test4
Fourth diagnostic test variable (optional).
- test4Positive
The positive level for Test 4.
- test5
Fifth diagnostic test variable (optional).
- test5Positive
The positive level for Test 5.
- method
Method for analyzing tests without gold standard.
- bootstrap
Calculate bootstrap confidence intervals.
- nboot
Number of bootstrap samples for confidence intervals.
- alpha
Alpha level for confidence intervals.
- verbose
Show detailed progress messages during bootstrap analysis.
Value
A results object containing:
results$instructions | a html | ||||
results$agreement_stats | a table | ||||
results$clinical_summary | a html | ||||
results$method_guide | a html | ||||
results$prevalence | a table | ||||
results$test_metrics | a table | ||||
results$model_fit | a table | ||||
results$crosstab | a table | ||||
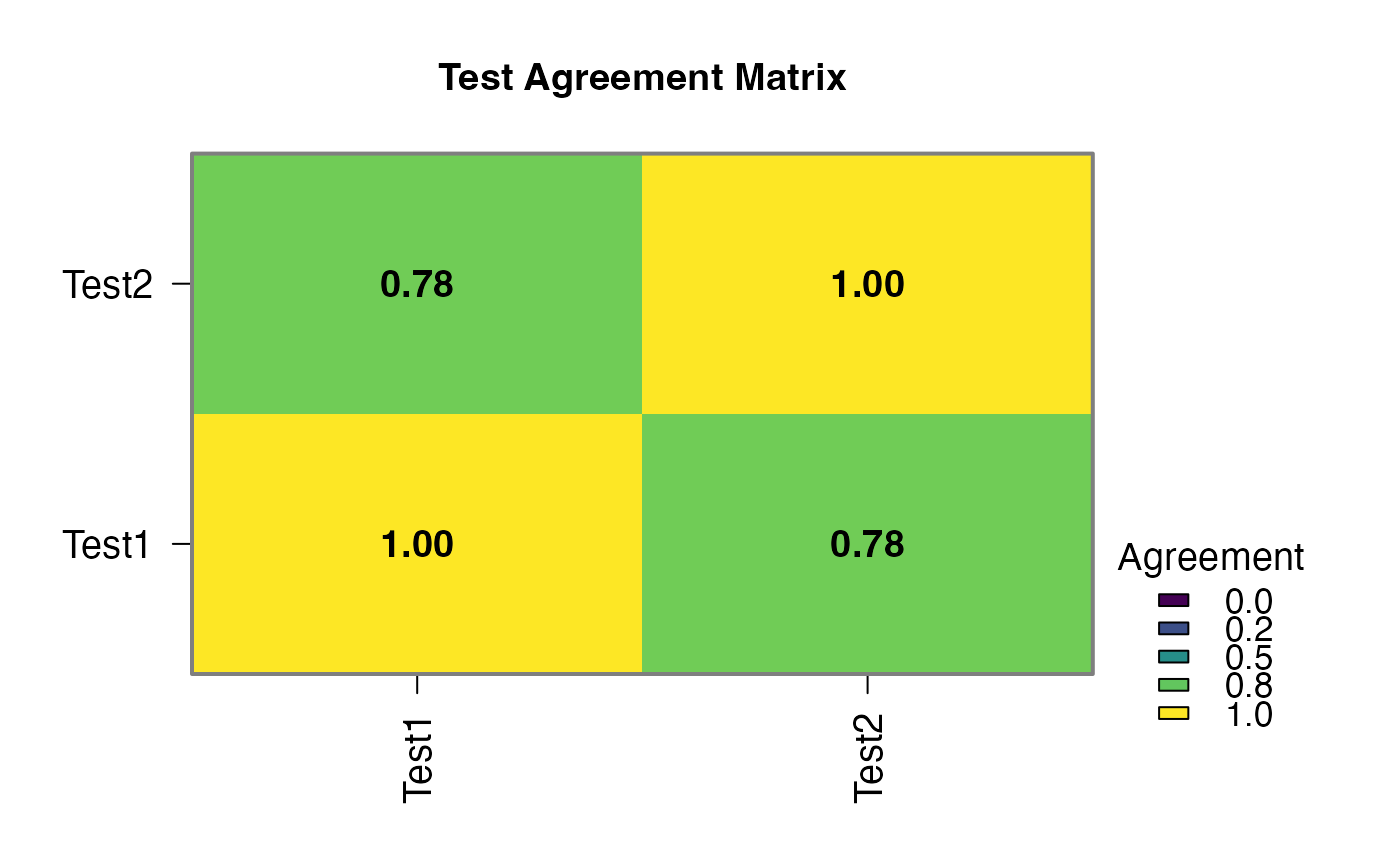
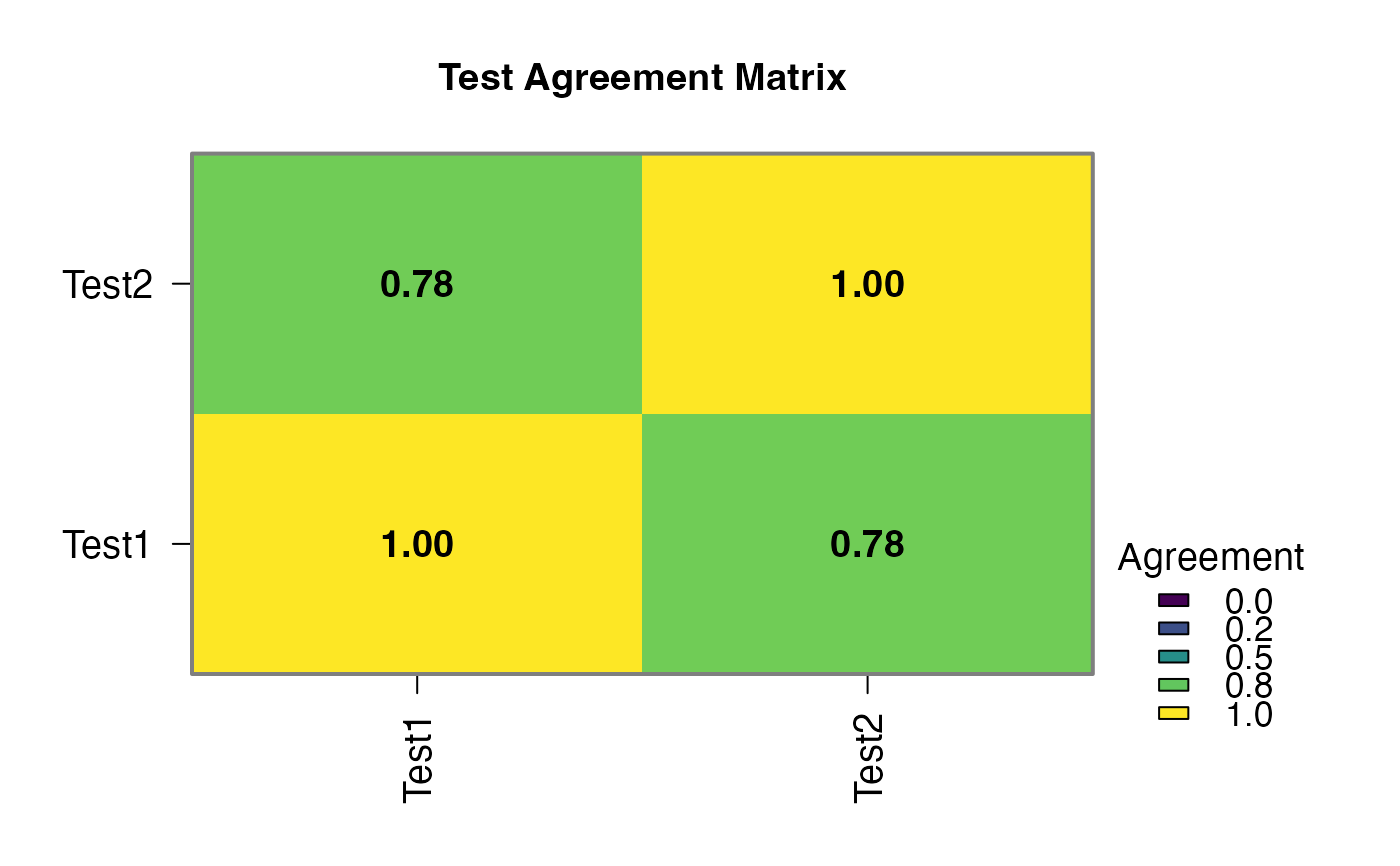
results$agreement_plot | an image | ||||
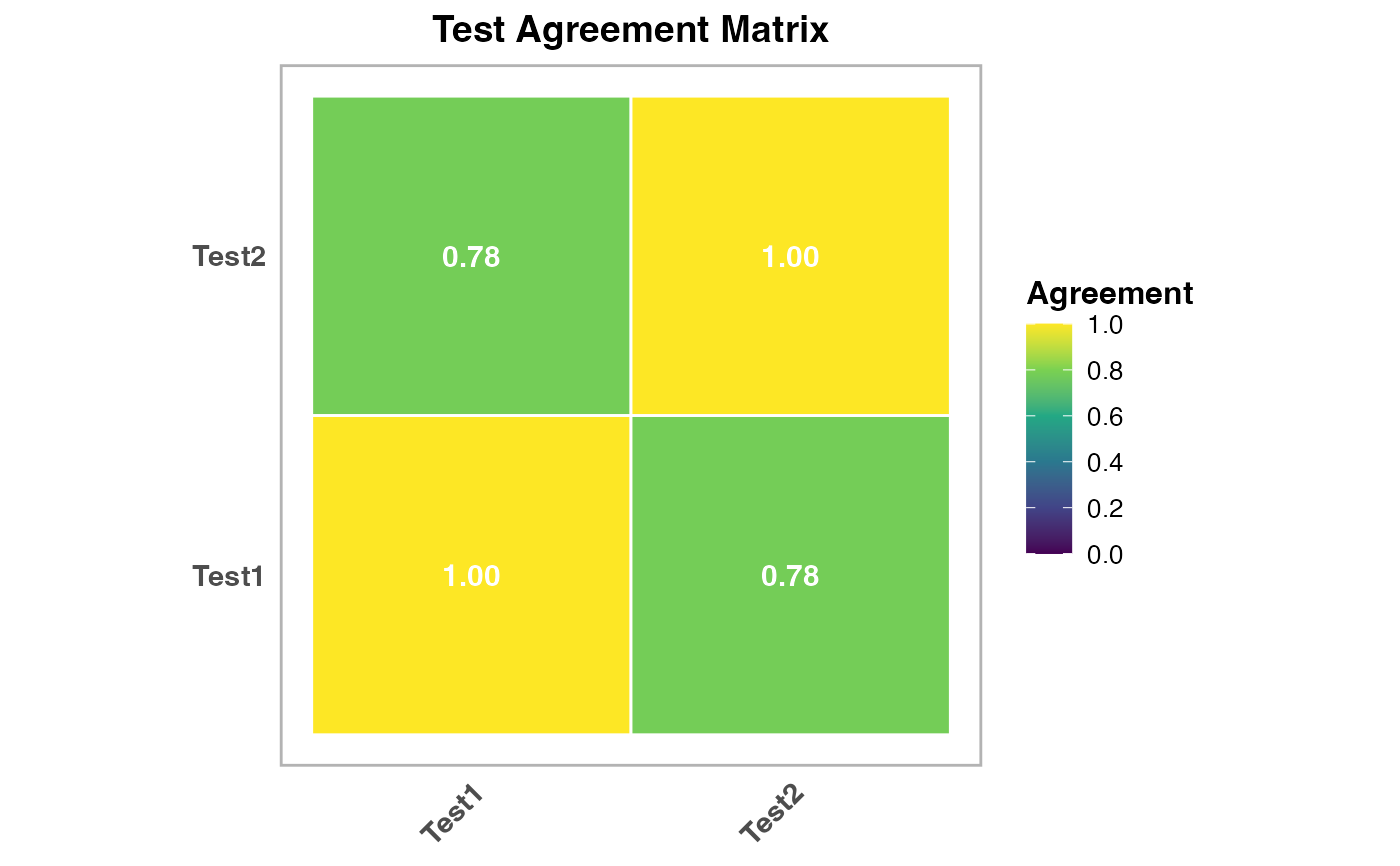
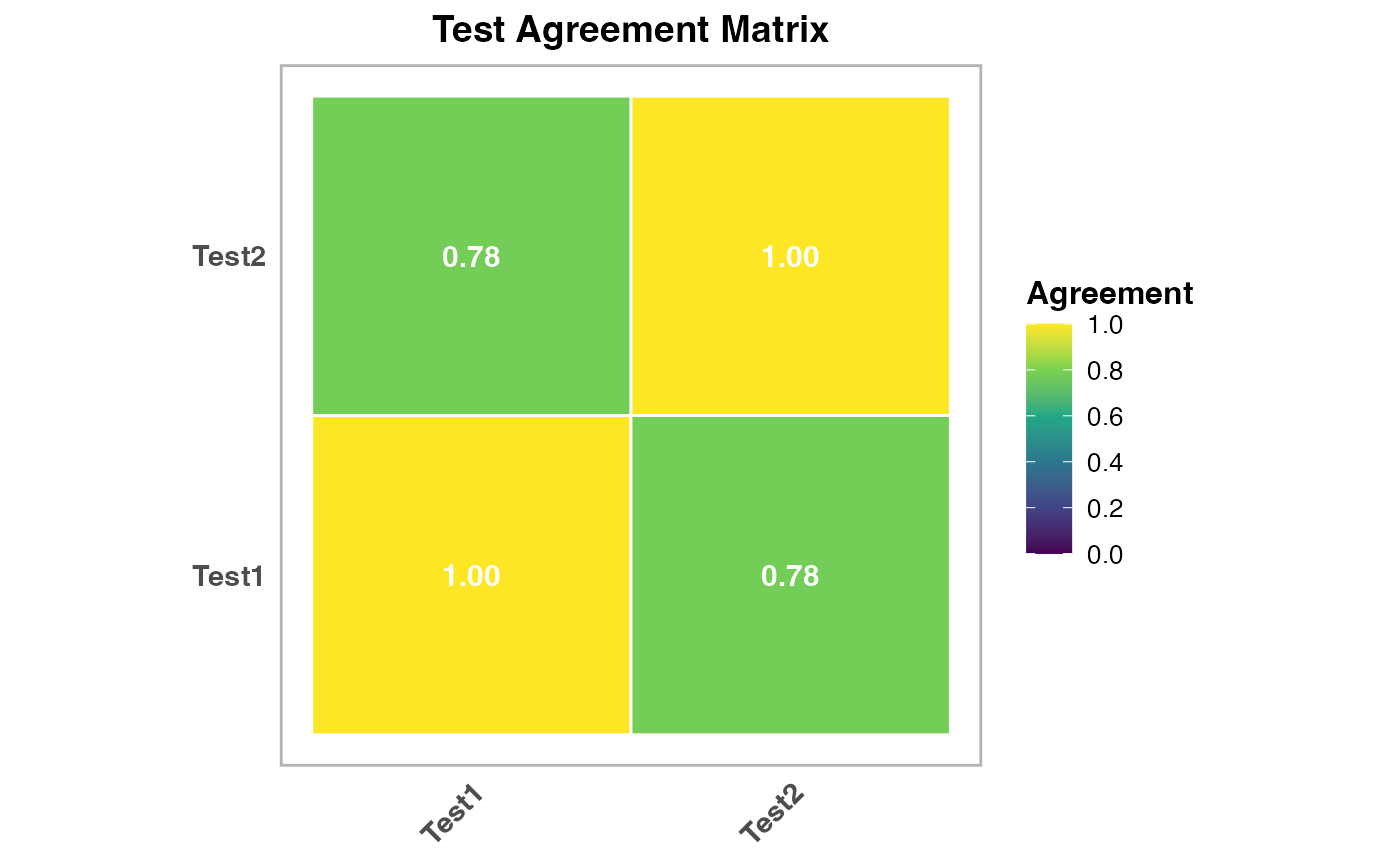
results$agreement_plot2 | an image |
Tables can be converted to data frames with asDF or as.data.frame. For example:
results$agreement_stats$asDF
as.data.frame(results$agreement_stats)
Examples
# \donttest{
# Basic example with simulated diagnostic test data
set.seed(123)
n <- 200
# Simulate disease status (latent, unknown)
disease <- rbinom(n, 1, 0.3) # 30 percent prevalence
# Simulate test results with known sensitivity/specificity
test1_result <- ifelse(disease == 1,
rbinom(sum(disease), 1, 0.85), # sensitivity 0.85
rbinom(sum(1-disease), 1, 0.15)) # 1-specificity 0.15
test1_result <- factor(test1_result, levels=c(0,1), labels=c("Negative", "Positive"))
test2_result <- ifelse(disease == 1,
rbinom(sum(disease), 1, 0.80), # sensitivity 0.80
rbinom(sum(1-disease), 1, 0.10)) # 1-specificity 0.10
test2_result <- factor(test2_result, levels=c(0,1), labels=c("Negative", "Positive"))
# Create data frame
test_data <- data.frame(
Test1 = test1_result,
Test2 = test2_result
)
# Latent Class Analysis (recommended method)
nogoldstandard(
data = test_data,
test1 = "Test1",
test1Positive = "Positive",
test2 = "Test2",
test2Positive = "Positive",
test3Positive = "Positive",
test4Positive = "Positive",
test5Positive = "Positive",
method = "composite"
)
#>
#> ANALYSIS WITHOUT GOLD STANDARD
#>
#> Composite reference with even number of tests may result in ties.
#> Consider using an odd number of tests or a different method.
#>
#> Agreement Statistics (Cohen's Kappa)
#> ──────────────────────────────────────────────────────────
#> Test Pair Kappa p-value Agreement
#> ──────────────────────────────────────────────────────────
#> Test1 vs Test2 0.5038900 < .0000001 78.00000
#> ──────────────────────────────────────────────────────────
#>
#>
#> <div class='clinical-summary' style='background: #f0f8ff; padding:
#> 15px; border-radius: 8px; margin: 10px 0;'><h4 style='color: #1565c0;
#> margin-top: 0;'>📋 Clinical Summary
#>
#> Analysis: No gold standard analysis using composite method
#>
#> Tests analyzed: Test1, Test2 (N=2)
#>
#> Disease prevalence: 44.0%
#>
#> Test sensitivities: Range from 69.3% to 80.7%
#>
#> Clinical interpretation: High prevalence setting - high PPV expected,
#> focus on confirming disease
#>
#> <div style='background: #f8f9fa; padding: 20px; border-radius: 8px;
#> margin: 15px 0; border-left: 4px solid #007bff;'><h3 style='color:
#> #007bff; margin-top: 0;'>📖 Method Selection Guide
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #2e7d32; margin-top: 0;'>🏆
#> Latent Class Analysis (Recommended)
#>
#> Description: Most robust method using mixture models. Estimates
#> disease prevalence and test parameters simultaneously.
#>
#> Best for: Diagnostic validation studies with 3+ tests and N≥100
#>
#> Strengths: Handles conditional dependence, provides model fit
#> statistics, most statistically rigorous
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e3f2fd;
#> border-radius: 5px;'><h4 style='color: #1565c0; margin-top: 0;'>📊
#> Bayesian Analysis
#>
#> Description: Incorporates prior knowledge about test performance using
#> Bayesian methods.
#>
#> Best for: Studies where you have prior information about expected
#> sensitivity/specificity
#>
#> Strengths: Uses prior knowledge, handles uncertainty well, good for
#> smaller samples
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fff3e0;
#> border-radius: 5px;'><h4 style='color: #ef6c00; margin-top: 0;'>🗳️
#> Composite Reference
#>
#> Description: Uses majority vote of available tests as pseudo-gold
#> standard.
#>
#> Best for: Inter-rater agreement studies with 3+ tests, exploratory
#> analysis
#>
#> Strengths: Simple and intuitive, requires minimal assumptions, good
#> starting point
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fce4ec;
#> border-radius: 5px;'><h4 style='color: #c2185b; margin-top: 0;'>🔒 All
#> Tests Positive
#>
#> Description: Conservative approach - disease present only if ALL tests
#> are positive.
#>
#> Best for: Highly specific diagnoses where false positives are very
#> costly
#>
#> Strengths: High specificity reference, minimizes false positives
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #388e3c; margin-top: 0;'>🔓 Any
#> Test Positive
#>
#> Description: Liberal approach - disease present if ANY test is
#> positive.
#>
#> Best for: Population screening scenarios where missing cases is costly
#>
#> Strengths: High sensitivity reference, minimizes false negatives
#>
#> <div style='margin: 15px 0; padding: 10px; background: #fff8e1;
#> border-radius: 5px; border-left: 3px solid #ffb300;'><h4 style='color:
#> #e65100; margin-top: 0;'>💡 Selection Tips
#>
#> Start with Latent Class Analysis for most diagnostic studiesUse
#> Composite Reference for quick exploratory analysisChoose All/Any Tests
#> Positive based on clinical consequences of errorsConsider Bayesian if
#> you have strong prior information
#>
#> Disease Prevalence
#> ───────────────────────────────────────
#> Estimate Lower CI Upper CI
#> ───────────────────────────────────────
#> 44.00000 37.12055 50.87945
#> ───────────────────────────────────────
#>
#>
#> Test Performance Metrics
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test Sensitivity Lower CI Upper CI Specificity Lower CI Upper CI PPV NPV
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test1 80.68182 75.21035 86.15329 100.00000 100.00000 100.00000 100.00000 86.82171
#> Test2 69.31818 62.92676 75.70960 100.00000 100.00000 100.00000 100.00000 80.57554
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Test Cross-Tabulation
#> ───────────────────────────────────────────
#> Test Combination Count Percentage
#> ───────────────────────────────────────────
#> Test1-, Test2- 112 56.00000
#> Test1+, Test2+ 44 22.00000
#> Test1+, Test2- 27 13.50000
#> Test1-, Test2+ 17 8.50000
#> ───────────────────────────────────────────
#>
 # With bootstrap confidence intervals
nogoldstandard(
data = test_data,
test1 = "Test1",
test1Positive = "Positive",
test2 = "Test2",
test2Positive = "Positive",
test3Positive = "Positive",
test4Positive = "Positive",
test5Positive = "Positive",
method = "composite",
bootstrap = TRUE,
nboot = 500,
verbose = TRUE
)
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for prevalence
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.2 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5023.52 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for sensitivity
#> Test index: 1
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5662.19 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for specificity
#> Test index: 1
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (6071.50 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for sensitivity
#> Test index: 2
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.1 sec elapsed, ~0.1 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5985.38 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for specificity
#> Test index: 2
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (6891.14 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> ANALYSIS WITHOUT GOLD STANDARD
#>
#> Composite reference with even number of tests may result in ties.
#> Consider using an odd number of tests or a different method.
#>
#> Clinical validation: 2 tests analyzed with N=200 using composite
#> method
#>
#> Agreement Statistics (Cohen's Kappa)
#> ──────────────────────────────────────────────────────────
#> Test Pair Kappa p-value Agreement
#> ──────────────────────────────────────────────────────────
#> Test1 vs Test2 0.5038900 < .0000001 78.00000
#> ──────────────────────────────────────────────────────────
#>
#>
#> <div class='clinical-summary' style='background: #f0f8ff; padding:
#> 15px; border-radius: 8px; margin: 10px 0;'><h4 style='color: #1565c0;
#> margin-top: 0;'>📋 Clinical Summary
#>
#> Analysis: No gold standard analysis using composite method
#>
#> Tests analyzed: Test1, Test2 (N=2)
#>
#> Disease prevalence: 44.0%
#>
#> Test sensitivities: Range from 69.3% to 80.7%
#>
#> Clinical interpretation: High prevalence setting - high PPV expected,
#> focus on confirming disease
#>
#> <div style='background: #f8f9fa; padding: 20px; border-radius: 8px;
#> margin: 15px 0; border-left: 4px solid #007bff;'><h3 style='color:
#> #007bff; margin-top: 0;'>📖 Method Selection Guide
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #2e7d32; margin-top: 0;'>🏆
#> Latent Class Analysis (Recommended)
#>
#> Description: Most robust method using mixture models. Estimates
#> disease prevalence and test parameters simultaneously.
#>
#> Best for: Diagnostic validation studies with 3+ tests and N≥100
#>
#> Strengths: Handles conditional dependence, provides model fit
#> statistics, most statistically rigorous
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e3f2fd;
#> border-radius: 5px;'><h4 style='color: #1565c0; margin-top: 0;'>📊
#> Bayesian Analysis
#>
#> Description: Incorporates prior knowledge about test performance using
#> Bayesian methods.
#>
#> Best for: Studies where you have prior information about expected
#> sensitivity/specificity
#>
#> Strengths: Uses prior knowledge, handles uncertainty well, good for
#> smaller samples
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fff3e0;
#> border-radius: 5px;'><h4 style='color: #ef6c00; margin-top: 0;'>🗳️
#> Composite Reference
#>
#> Description: Uses majority vote of available tests as pseudo-gold
#> standard.
#>
#> Best for: Inter-rater agreement studies with 3+ tests, exploratory
#> analysis
#>
#> Strengths: Simple and intuitive, requires minimal assumptions, good
#> starting point
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fce4ec;
#> border-radius: 5px;'><h4 style='color: #c2185b; margin-top: 0;'>🔒 All
#> Tests Positive
#>
#> Description: Conservative approach - disease present only if ALL tests
#> are positive.
#>
#> Best for: Highly specific diagnoses where false positives are very
#> costly
#>
#> Strengths: High specificity reference, minimizes false positives
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #388e3c; margin-top: 0;'>🔓 Any
#> Test Positive
#>
#> Description: Liberal approach - disease present if ANY test is
#> positive.
#>
#> Best for: Population screening scenarios where missing cases is costly
#>
#> Strengths: High sensitivity reference, minimizes false negatives
#>
#> <div style='margin: 15px 0; padding: 10px; background: #fff8e1;
#> border-radius: 5px; border-left: 3px solid #ffb300;'><h4 style='color:
#> #e65100; margin-top: 0;'>💡 Selection Tips
#>
#> Start with Latent Class Analysis for most diagnostic studiesUse
#> Composite Reference for quick exploratory analysisChoose All/Any Tests
#> Positive based on clinical consequences of errorsConsider Bayesian if
#> you have strong prior information
#>
#> Disease Prevalence
#> ───────────────────────────────────────
#> Estimate Lower CI Upper CI
#> ───────────────────────────────────────
#> 44.00000 37.00000 51.00000
#> ───────────────────────────────────────
#>
#>
#> Test Performance Metrics
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test Sensitivity Lower CI Upper CI Specificity Lower CI Upper CI PPV NPV
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test1 80.68182 72.56270 87.95732 100.00000 100.00000 100.00000 100.00000 86.82171
#> Test2 69.31818 59.23367 78.44685 100.00000 100.00000 100.00000 100.00000 80.57554
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Test Cross-Tabulation
#> ───────────────────────────────────────────
#> Test Combination Count Percentage
#> ───────────────────────────────────────────
#> Test1-, Test2- 112 56.00000
#> Test1+, Test2+ 44 22.00000
#> Test1+, Test2- 27 13.50000
#> Test1-, Test2+ 17 8.50000
#> ───────────────────────────────────────────
#>
# With bootstrap confidence intervals
nogoldstandard(
data = test_data,
test1 = "Test1",
test1Positive = "Positive",
test2 = "Test2",
test2Positive = "Positive",
test3Positive = "Positive",
test4Positive = "Positive",
test5Positive = "Positive",
method = "composite",
bootstrap = TRUE,
nboot = 500,
verbose = TRUE
)
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for prevalence
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.2 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5023.52 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for sensitivity
#> Test index: 1
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5662.19 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for specificity
#> Test index: 1
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (6071.50 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for sensitivity
#> Test index: 2
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.1 sec elapsed, ~0.1 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (5985.38 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> === Bootstrap Analysis ===
#> Starting bootstrap with 500 iterations for composite method
#> Estimating confidence intervals for specificity
#> Test index: 2
#> 25/500 (5.0%) - 25 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 50/500 (10.0%) - 50 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 75/500 (15.0%) - 75 successful, 0 errors - 0.0 sec elapsed, ~0.1 sec remaining
#> 100/500 (20.0%) - 100 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 125/500 (25.0%) - 125 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 150/500 (30.0%) - 150 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 175/500 (35.0%) - 175 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 200/500 (40.0%) - 200 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 225/500 (45.0%) - 225 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 250/500 (50.0%) - 250 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 275/500 (55.0%) - 275 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 300/500 (60.0%) - 300 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 325/500 (65.0%) - 325 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 350/500 (70.0%) - 350 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 375/500 (75.0%) - 375 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 400/500 (80.0%) - 400 successful, 0 errors - 0.0 sec elapsed, ~0.0 sec remaining
#> 425/500 (85.0%) - 425 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 450/500 (90.0%) - 450 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 475/500 (95.0%) - 475 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#> 500/500 (100.0%) - 500 successful, 0 errors - 0.1 sec elapsed, ~0.0 sec remaining
#>
#> === Bootstrap Complete ===
#> Total time: 0.1 seconds (6891.14 iterations/sec)
#> Successful iterations: 500 (100.0%)
#> Failed iterations: 0 (0.0%)
#> Warning: 2 arguments not used by format 'Confidence interval (%.1f%%):'
#> Confidence interval (95.0%):
#>
#> ANALYSIS WITHOUT GOLD STANDARD
#>
#> Composite reference with even number of tests may result in ties.
#> Consider using an odd number of tests or a different method.
#>
#> Clinical validation: 2 tests analyzed with N=200 using composite
#> method
#>
#> Agreement Statistics (Cohen's Kappa)
#> ──────────────────────────────────────────────────────────
#> Test Pair Kappa p-value Agreement
#> ──────────────────────────────────────────────────────────
#> Test1 vs Test2 0.5038900 < .0000001 78.00000
#> ──────────────────────────────────────────────────────────
#>
#>
#> <div class='clinical-summary' style='background: #f0f8ff; padding:
#> 15px; border-radius: 8px; margin: 10px 0;'><h4 style='color: #1565c0;
#> margin-top: 0;'>📋 Clinical Summary
#>
#> Analysis: No gold standard analysis using composite method
#>
#> Tests analyzed: Test1, Test2 (N=2)
#>
#> Disease prevalence: 44.0%
#>
#> Test sensitivities: Range from 69.3% to 80.7%
#>
#> Clinical interpretation: High prevalence setting - high PPV expected,
#> focus on confirming disease
#>
#> <div style='background: #f8f9fa; padding: 20px; border-radius: 8px;
#> margin: 15px 0; border-left: 4px solid #007bff;'><h3 style='color:
#> #007bff; margin-top: 0;'>📖 Method Selection Guide
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #2e7d32; margin-top: 0;'>🏆
#> Latent Class Analysis (Recommended)
#>
#> Description: Most robust method using mixture models. Estimates
#> disease prevalence and test parameters simultaneously.
#>
#> Best for: Diagnostic validation studies with 3+ tests and N≥100
#>
#> Strengths: Handles conditional dependence, provides model fit
#> statistics, most statistically rigorous
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e3f2fd;
#> border-radius: 5px;'><h4 style='color: #1565c0; margin-top: 0;'>📊
#> Bayesian Analysis
#>
#> Description: Incorporates prior knowledge about test performance using
#> Bayesian methods.
#>
#> Best for: Studies where you have prior information about expected
#> sensitivity/specificity
#>
#> Strengths: Uses prior knowledge, handles uncertainty well, good for
#> smaller samples
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fff3e0;
#> border-radius: 5px;'><h4 style='color: #ef6c00; margin-top: 0;'>🗳️
#> Composite Reference
#>
#> Description: Uses majority vote of available tests as pseudo-gold
#> standard.
#>
#> Best for: Inter-rater agreement studies with 3+ tests, exploratory
#> analysis
#>
#> Strengths: Simple and intuitive, requires minimal assumptions, good
#> starting point
#>
#> <div style='margin: 15px 0; padding: 15px; background: #fce4ec;
#> border-radius: 5px;'><h4 style='color: #c2185b; margin-top: 0;'>🔒 All
#> Tests Positive
#>
#> Description: Conservative approach - disease present only if ALL tests
#> are positive.
#>
#> Best for: Highly specific diagnoses where false positives are very
#> costly
#>
#> Strengths: High specificity reference, minimizes false positives
#>
#> <div style='margin: 15px 0; padding: 15px; background: #e8f5e8;
#> border-radius: 5px;'><h4 style='color: #388e3c; margin-top: 0;'>🔓 Any
#> Test Positive
#>
#> Description: Liberal approach - disease present if ANY test is
#> positive.
#>
#> Best for: Population screening scenarios where missing cases is costly
#>
#> Strengths: High sensitivity reference, minimizes false negatives
#>
#> <div style='margin: 15px 0; padding: 10px; background: #fff8e1;
#> border-radius: 5px; border-left: 3px solid #ffb300;'><h4 style='color:
#> #e65100; margin-top: 0;'>💡 Selection Tips
#>
#> Start with Latent Class Analysis for most diagnostic studiesUse
#> Composite Reference for quick exploratory analysisChoose All/Any Tests
#> Positive based on clinical consequences of errorsConsider Bayesian if
#> you have strong prior information
#>
#> Disease Prevalence
#> ───────────────────────────────────────
#> Estimate Lower CI Upper CI
#> ───────────────────────────────────────
#> 44.00000 37.00000 51.00000
#> ───────────────────────────────────────
#>
#>
#> Test Performance Metrics
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test Sensitivity Lower CI Upper CI Specificity Lower CI Upper CI PPV NPV
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Test1 80.68182 72.56270 87.95732 100.00000 100.00000 100.00000 100.00000 86.82171
#> Test2 69.31818 59.23367 78.44685 100.00000 100.00000 100.00000 100.00000 80.57554
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Test Cross-Tabulation
#> ───────────────────────────────────────────
#> Test Combination Count Percentage
#> ───────────────────────────────────────────
#> Test1-, Test2- 112 56.00000
#> Test1+, Test2+ 44 22.00000
#> Test1+, Test2- 27 13.50000
#> Test1-, Test2+ 17 8.50000
#> ───────────────────────────────────────────
#>
 # }
# }