Conditional Survival Estimation - Comprehensive Guide
ClinicoPath Development Team
2026-03-22
Source:vignettes/jsurvival-conditionalsurvival-comprehensive.Rmd
jsurvival-conditionalsurvival-comprehensive.RmdNote:
conditionalsurvival()is designed primarily for the jamovi GUI. The R syntax shown here is for reference and advanced scripting.
Conditional Survival Estimation
Overview
In clinical oncology, a patient’s prognosis improves with each additional year of disease-free survival. Standard Kaplan-Meier estimates describe the probability of surviving beyond time t from the moment of diagnosis or treatment. Conditional survival answers a different and often more useful question: given that a patient has already survived to time s, what is their probability of surviving to time t?
Formally: CS(t | s) = P(T > t | T > s) = S(t) / S(s)
This is implemented via the conditionalsurvival()
function in the jsurvival module (menu: SurvivalD > ClinicoPath
Survival). Four estimation methods are available:
| Method | Key | Description |
|---|---|---|
| Kaplan-Meier Weights | km |
Weighted KM estimation via condSURV::KMW() (falls back
to S(t)/S(s) ratio when condSURV is not available) |
| Landmark Approach | landmark |
Subsets cohort to patients surviving past the conditioning time, then refits KM |
| Inverse Probability Weighting | ipw |
IPCW-based estimation (stub; currently falls back to manual KM ratio) |
| Presmoothed Kaplan-Meier | pkm |
Kernel-smoothed KM (stub; currently falls back to manual KM ratio) |
Datasets
This vignette uses the bundled conditionalsurvival_test
dataset:
| Dataset | N | Scenario | Key Variables |
|---|---|---|---|
conditionalsurvival_test |
150 | Colorectal cancer, Weibull survival, stage-dependent hazard |
OverallTime (months), Event (0/1),
EventFactor (Alive/Dead), Treatment,
Stage, Grade, Age,
Sex, CEA
|
histopathology |
varies | Built-in ClinicoPath dataset | General clinicopathological variables |
data("conditionalsurvival_test")
str(conditionalsurvival_test)
#> 'data.frame': 150 obs. of 10 variables:
#> $ PatientID : chr "CS-001" "CS-002" "CS-003" "CS-004" ...
#> $ OverallTime: num 10.6 70.4 31.6 18.7 28.4 10.7 5 16.1 20.5 35 ...
#> $ Event : int 1 1 0 1 1 1 1 1 1 0 ...
#> $ EventFactor: Factor w/ 2 levels "Alive","Dead": 2 2 1 2 2 2 2 2 2 1 ...
#> $ Age : num 69 50 65 62 55 33 54 51 64 57 ...
#> $ Sex : Factor w/ 2 levels "Female","Male": 2 2 1 2 1 2 1 2 1 1 ...
#> $ Treatment : Factor w/ 2 levels "Surgery","Surgery+Chemo": 1 2 2 1 2 2 1 1 1 1 ...
#> $ Stage : Factor w/ 4 levels "I","II","III",..: 2 2 2 4 3 3 4 2 3 1 ...
#> $ Grade : Factor w/ 3 levels "Well","Moderate",..: 1 2 1 3 2 3 3 2 3 1 ...
#> $ CEA : num 13.1 15.5 12.2 17.2 20.3 16.9 19.5 15.3 15.4 20.6 ...
summary(conditionalsurvival_test[, c("OverallTime", "Event", "Treatment", "Stage")])
#> OverallTime Event Treatment Stage
#> Min. : 0.70 Min. :0.0 Surgery :67 I :26
#> 1st Qu.: 10.80 1st Qu.:1.0 Surgery+Chemo:83 II :43
#> Median : 24.85 Median :1.0 III:49
#> Mean : 30.36 Mean :0.8 IV :32
#> 3rd Qu.: 44.70 3rd Qu.:1.0
#> Max. :117.80 Max. :1.0Basic Usage
Unstratified Analysis (Overall Cohort)
The simplest use case: estimate conditional survival for the entire
cohort with default settings. When conditionTime = 0, the
function automatically uses the median follow-up time.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 0,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = TRUE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultUnstratified with Fixed Conditioning Time
Set conditionTime = 12 to ask: “Given a patient survived
12 months, what is their probability of surviving to each subsequent
time point?”
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultStratified Analysis
By Treatment
Compare conditional survival between Surgery and Surgery+Chemo
groups. The conditionVar option activates stratified
analysis, producing separate conditional survival estimates and curves
per group.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = TRUE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Method: Kaplan-Meier Weights
#>
#> Analysis Summary:
#>
#> Conditioning Time: 12Number of Time Points: 10Average Conditional
#> Survival: 0.239Range: 0.031 - 0.626
#>
#> Clinical Interpretation:
#>
#> These conditional survival probabilities represent updated survival
#> estimates for patients who have survived to time 12. For example, a
#> conditional survival probability of 0.80 at time point 117.8 means
#> that patients who have survived to time 12 have an 80% probability of
#> surviving to time 117.8.
#>
#> Method Description:
#>
#> Kaplan-Meier Weights Method: This approach uses Kaplan-Meier weights
#> to estimate conditional survival probabilities. The weights account
#> for the censoring pattern and provide more accurate estimates when
#> censoring is informative.
#>
#> Statistical Notes:
#>
#> Confidence intervals are calculated using the specified confidence
#> level (95%)Standard errors are computed using appropriate methods for
#> each estimation approachTime points at or before the conditioning time
#> have conditional survival probability = 1.0
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the Surgery group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 3.1% (95% CI:
#> 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Surgery+Chemo group (n
#> at risk = 110), the estimated conditional 117.8-unit survival
#> probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').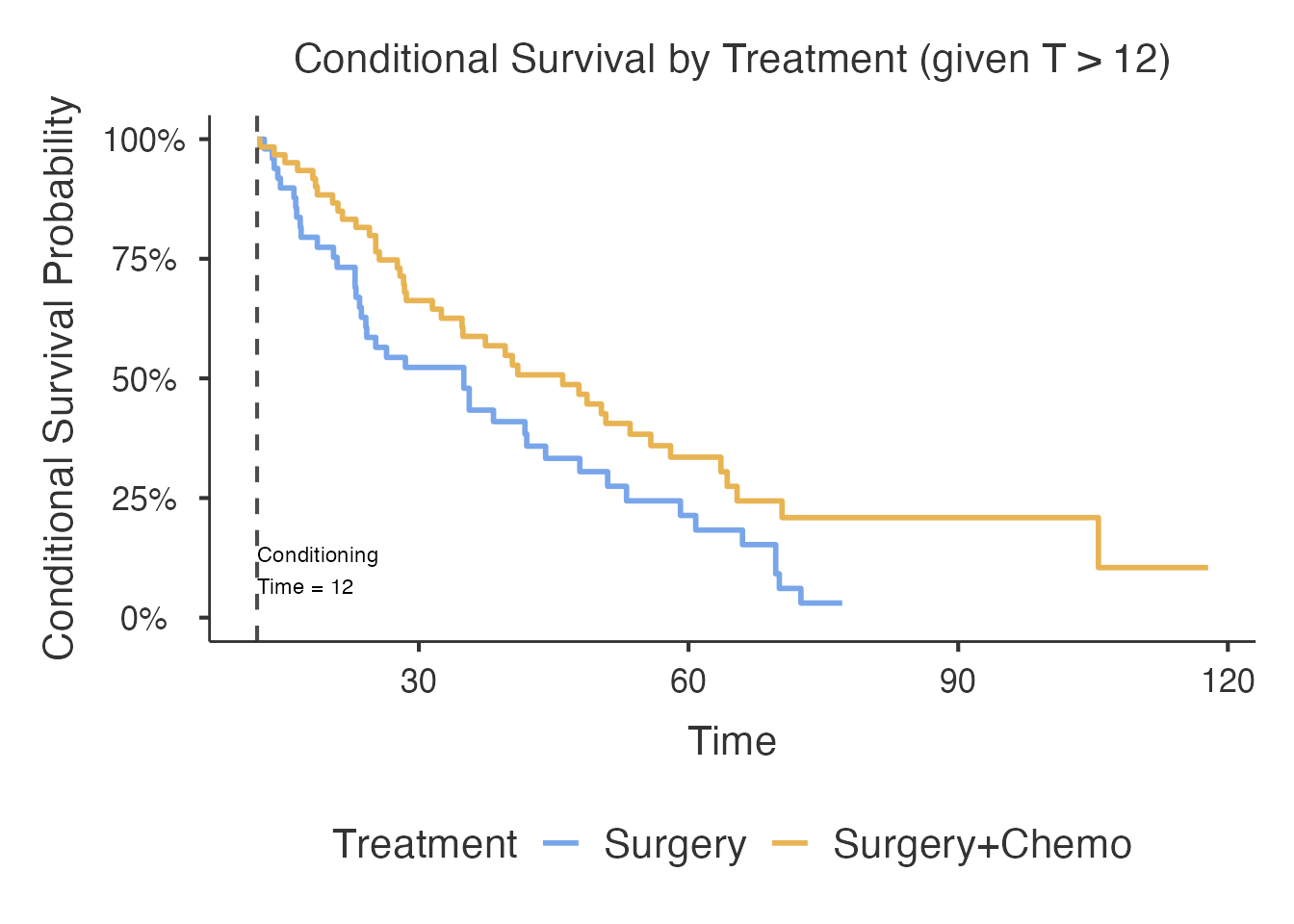
By Stage
Stage has 4 levels (I, II, III, IV). Groups with fewer than 3 events are silently skipped. Early-stage patients with few events at late conditioning times may be excluded automatically.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Stage,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> I 33.2000000 12.0000000 0.81136364 ᵃ 0.12822125 0.560054614 1.0000000
#> I 54.3000000 12.0000000 0.54793388 0.13922208 0.275063631 0.8208041
#> I 75.5000000 12.0000000 0.20547521 0.10733848 0.000000000 0.4158548
#> I 96.6000000 12.0000000 0.20547521 0.10733848 0.000000000 0.4158548
#> I 117.8000000 12.0000000 0.00000000 0.00000000 0.000000000 0.0000000
#> II 33.2000000 12.0000000 0.80557977 0.09624791 0.616937341 0.9942222
#> II 54.3000000 12.0000000 0.43366620 0.09857655 0.240459724 0.6268727
#> II 75.5000000 12.0000000 0.17346648 0.08556952 0.005753307 0.3411797
#> II 96.6000000 12.0000000 0.17346648 0.08556952 0.005753307 0.3411797
#> II 117.8000000 12.0000000 0.17346648 0.08556952 0.005753307 0.3411797
#> III 33.2000000 12.0000000 0.41940789 0.09143042 0.240207571 0.5986082
#> III 54.3000000 12.0000000 0.21303258 0.07335106 0.069267153 0.3567980
#> III 75.5000000 12.0000000 0.05680869 0.05191773 0.000000000 0.1585656
#> III 96.6000000 12.0000000 0.05680869 0.05191773 0.000000000 0.1585656
#> III 117.8000000 12.0000000 0.05680869 0.05191773 0.000000000 0.1585656
#> IV 33.2000000 12.0000000 0.07692308 0.07727458 0.000000000 0.2283785
#> IV 54.3000000 12.0000000 0.00000000 0.00000000 0.000000000 0.0000000
#> IV 75.5000000 12.0000000 0.00000000 0.00000000 0.000000000 0.0000000
#> IV 96.6000000 12.0000000 0.00000000 0.00000000 0.000000000 0.0000000
#> IV 117.8000000 12.0000000 0.00000000 0.00000000 0.000000000 0.0000000
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the I group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 0.0% (95% CI:
#> 0.0%--0.0%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the II group (n at risk =
#> 110), the estimated conditional 117.8-unit survival probability was
#> 17.3% (95% CI: 0.6%--34.1%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the III group (n at risk =
#> 110), the estimated conditional 117.8-unit survival probability was
#> 5.7% (95% CI: 0.0%--15.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the IV group (n at risk =
#> 110), the estimated conditional 117.8-unit survival probability was
#> 0.0% (95% CI: 0.0%--0.0%).<p style='font-size:0.85em;
#> color:#666;'>Tip: Copy the text above directly into your manuscript
#> methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').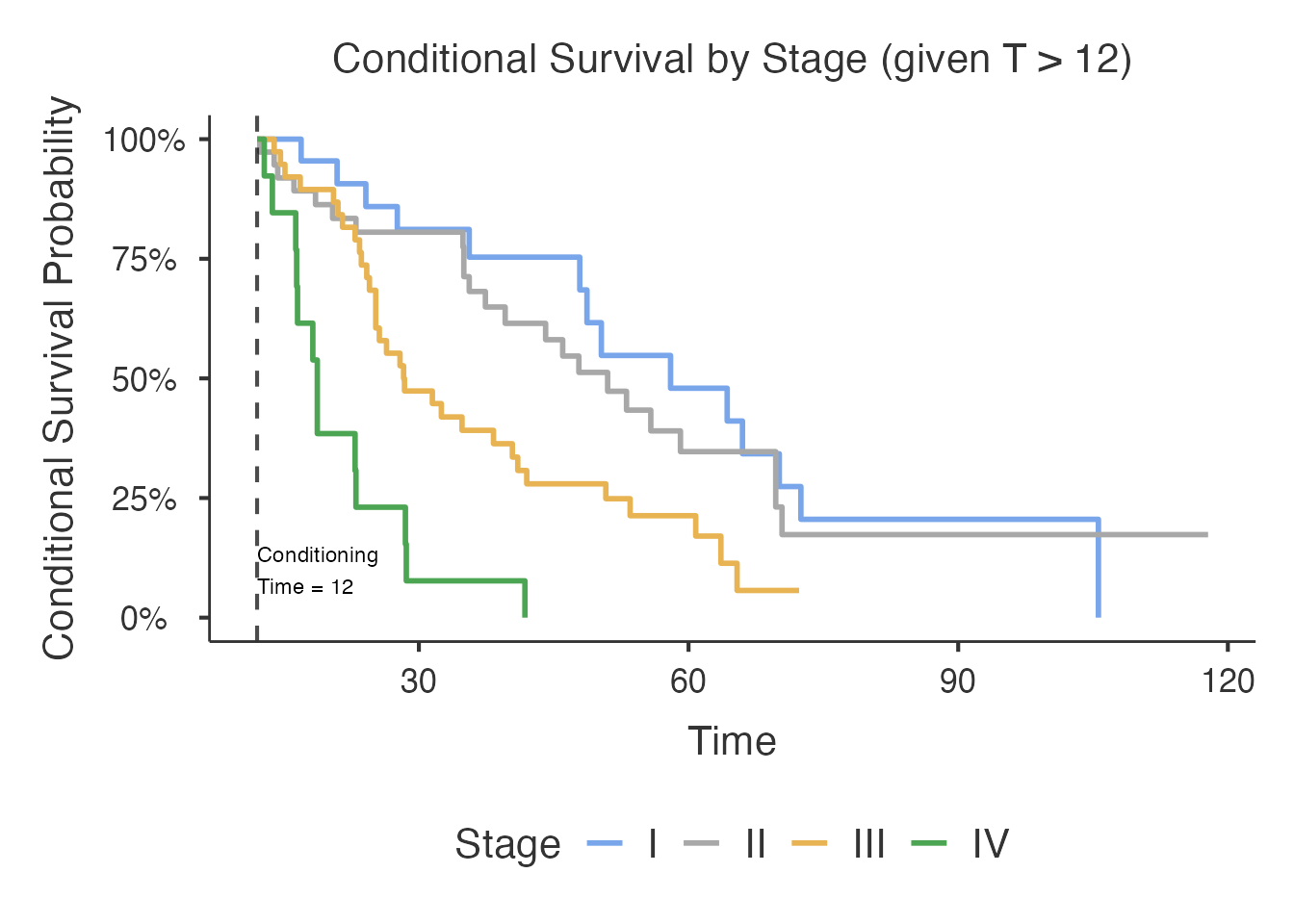
By Grade
Three tumor differentiation levels: Well, Moderate, Poor.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Grade,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Well 33.2000000 12.0000000 0.75574449 ᵃ 0.10345126 0.55298375 0.95850522
#> Well 54.3000000 12.0000000 0.50823722 0.10921171 0.29418620 0.72228823
#> Well 75.5000000 12.0000000 0.22588321 0.09717278 0.03542807 0.41633835
#> Well 96.6000000 12.0000000 0.22588321 0.09717278 0.03542807 0.41633835
#> Well 117.8000000 12.0000000 0.22588321 0.09717278 0.03542807 0.41633835
#> Moderate 33.2000000 12.0000000 0.60303288 0.09787145 0.41120836 0.79485740
#> Moderate 54.3000000 12.0000000 0.28770599 0.08148025 0.12800763 0.44740435
#> Moderate 75.5000000 12.0000000 0.15733921 0.07290934 0.01443954 0.30023889
#> Moderate 96.6000000 12.0000000 0.15733921 0.07290934 0.01443954 0.30023889
#> Moderate 117.8000000 12.0000000 0.07866961 0.06650862 0.00000000 0.20902411
#> Poor 33.2000000 12.0000000 0.38235294 0.10048060 0.18541459 0.57929129
#> Poor 54.3000000 12.0000000 0.20168067 0.07571703 0.05327803 0.35008332
#> Poor 75.5000000 12.0000000 0.03361345 0.03317838 0.00000000 0.09864188
#> Poor 96.6000000 12.0000000 0.03361345 0.03317838 0.00000000 0.09864188
#> Poor 117.8000000 12.0000000 0.03361345 0.03317838 0.00000000 0.09864188
#> ──────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the Well group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 22.6% (95% CI:
#> 3.5%--41.6%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Moderate group (n at
#> risk = 110), the estimated conditional 117.8-unit survival probability
#> was 7.9% (95% CI: 0.0%--20.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Poor group (n at risk =
#> 110), the estimated conditional 117.8-unit survival probability was
#> 3.4% (95% CI: 0.0%--9.9%).<p style='font-size:0.85em;
#> color:#666;'>Tip: Copy the text above directly into your manuscript
#> methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Estimation Methods
Kaplan-Meier Weights (Default)
The default method. When the condSURV package is
installed, uses KMW weights for estimation. Otherwise falls back to the
manual S(t)/S(s) ratio.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = TRUE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Method: Kaplan-Meier Weights
#>
#> Analysis Summary:
#>
#> Conditioning Time: 12Number of Time Points: 10Average Conditional
#> Survival: 0.239Range: 0.031 - 0.626
#>
#> Clinical Interpretation:
#>
#> These conditional survival probabilities represent updated survival
#> estimates for patients who have survived to time 12. For example, a
#> conditional survival probability of 0.80 at time point 117.8 means
#> that patients who have survived to time 12 have an 80% probability of
#> surviving to time 117.8.
#>
#> Method Description:
#>
#> Kaplan-Meier Weights Method: This approach uses Kaplan-Meier weights
#> to estimate conditional survival probabilities. The weights account
#> for the censoring pattern and provide more accurate estimates when
#> censoring is informative.
#>
#> Statistical Notes:
#>
#> Confidence intervals are calculated using the specified confidence
#> level (95%)Standard errors are computed using appropriate methods for
#> each estimation approachTime points at or before the conditioning time
#> have conditional survival probability = 1.0
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the Surgery group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 3.1% (95% CI:
#> 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Surgery+Chemo group (n
#> at risk = 110), the estimated conditional 117.8-unit survival
#> probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Landmark Approach
The landmark method subsets the data to only patients who survived past the conditioning time, then estimates survival from that point forward. This is the simplest and most transparent approach but discards information from patients censored before the landmark.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "landmark",
showTable = TRUE,
showPlot = TRUE,
showExplanations = TRUE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Method: Landmark Approach
#>
#> Analysis Summary:
#>
#> Conditioning Time: 12Number of Time Points: 10Average Conditional
#> Survival: 0.239Range: 0.031 - 0.626
#>
#> Clinical Interpretation:
#>
#> These conditional survival probabilities represent updated survival
#> estimates for patients who have survived to time 12. For example, a
#> conditional survival probability of 0.80 at time point 117.8 means
#> that patients who have survived to time 12 have an 80% probability of
#> surviving to time 117.8.
#>
#> Method Description:
#>
#> Landmark Approach: This method subsets the data to include only
#> patients who survived to the conditioning time point, then estimates
#> survival from that point forward. This approach is simple but may lose
#> information from patients censored before the landmark time.
#>
#> Statistical Notes:
#>
#> Confidence intervals are calculated using the specified confidence
#> level (95%)Standard errors are computed using appropriate methods for
#> each estimation approachTime points at or before the conditioning time
#> have conditional survival probability = 1.0
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the landmark method. Given survival to
#> 12.0 in the Surgery group (n at risk = 110), the estimated conditional
#> 117.8-unit survival probability was 3.1% (95% CI: 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the landmark method.
#> Given survival to 12.0 in the Surgery+Chemo group (n at risk = 110),
#> the estimated conditional 117.8-unit survival probability was 10.5%
#> (95% CI: 0.0%--26.4%).<p style='font-size:0.85em; color:#666;'>Tip:
#> Copy the text above directly into your manuscript methods/results
#> section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').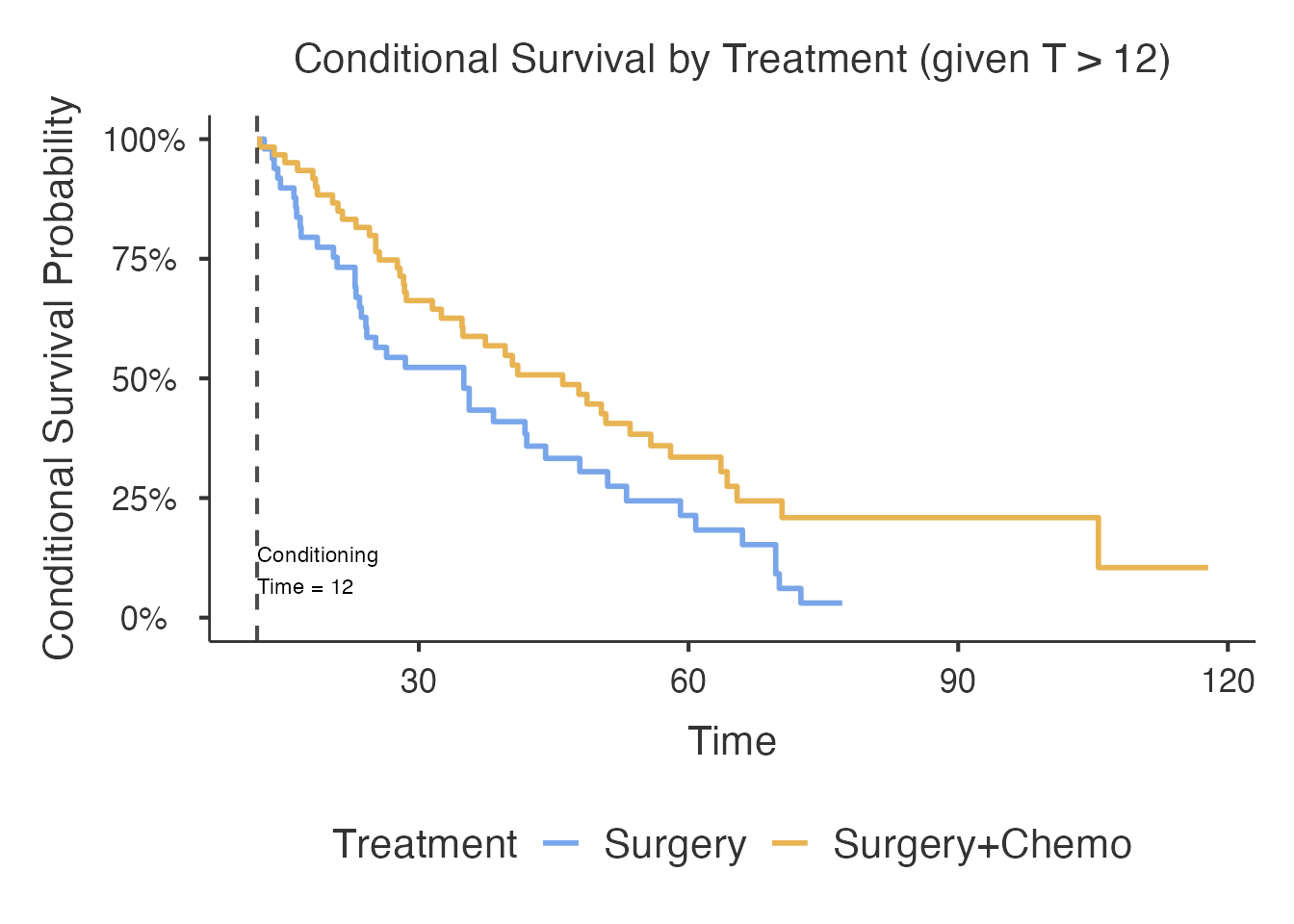
Inverse Probability Weighting
The IPW method is intended to use inverse probability of censoring weights for unbiased estimation. Currently implemented as a stub that falls back to the manual KM ratio method.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "ipw",
showTable = TRUE,
showPlot = FALSE,
showExplanations = TRUE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Method: Inverse Probability Weighting
#>
#> Analysis Summary:
#>
#> Conditioning Time: 12Number of Time Points: 10Average Conditional
#> Survival: 0.239Range: 0.031 - 0.626
#>
#> Clinical Interpretation:
#>
#> These conditional survival probabilities represent updated survival
#> estimates for patients who have survived to time 12. For example, a
#> conditional survival probability of 0.80 at time point 117.8 means
#> that patients who have survived to time 12 have an 80% probability of
#> surviving to time 117.8.
#>
#> Method Description:
#>
#> Inverse Probability Weighting: This approach uses inverse probability
#> weights to account for censoring, providing unbiased estimates of
#> conditional survival probabilities even when censoring is informative.
#>
#> Statistical Notes:
#>
#> Confidence intervals are calculated using the specified confidence
#> level (95%)Standard errors are computed using appropriate methods for
#> each estimation approachTime points at or before the conditioning time
#> have conditional survival probability = 1.0
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the inverse probability weighting method.
#> Given survival to 12.0 in the Surgery group (n at risk = 110), the
#> estimated conditional 117.8-unit survival probability was 3.1% (95%
#> CI: 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the inverse
#> probability weighting method. Given survival to 12.0 in the
#> Surgery+Chemo group (n at risk = 110), the estimated conditional
#> 117.8-unit survival probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Presmoothed Kaplan-Meier
The PKM method applies kernel smoothing to the KM estimator.
Currently a stub that falls back to the manual KM ratio. The
bandwidth option is reserved for controlling the smoothing
bandwidth when this method is fully implemented.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "pkm",
bandwidth = 0,
showTable = TRUE,
showPlot = FALSE,
showExplanations = TRUE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Method: Presmoothed Kaplan-Meier
#>
#> Analysis Summary:
#>
#> Conditioning Time: 12Number of Time Points: 10Average Conditional
#> Survival: 0.239Range: 0.031 - 0.626
#>
#> Clinical Interpretation:
#>
#> These conditional survival probabilities represent updated survival
#> estimates for patients who have survived to time 12. For example, a
#> conditional survival probability of 0.80 at time point 117.8 means
#> that patients who have survived to time 12 have an 80% probability of
#> surviving to time 117.8.
#>
#> Method Description:
#>
#> Presmoothed Kaplan-Meier: This method applies smoothing techniques to
#> the Kaplan-Meier estimator to reduce variability and provide more
#> stable conditional survival estimates.
#>
#> Statistical Notes:
#>
#> Confidence intervals are calculated using the specified confidence
#> level (95%)Standard errors are computed using appropriate methods for
#> each estimation approachTime points at or before the conditioning time
#> have conditional survival probability = 1.0
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the presmoothed Kaplan-Meier method.
#> Given survival to 12.0 in the Surgery group (n at risk = 110), the
#> estimated conditional 117.8-unit survival probability was 3.1% (95%
#> CI: 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the presmoothed
#> Kaplan-Meier method. Given survival to 12.0 in the Surgery+Chemo group
#> (n at risk = 110), the estimated conditional 117.8-unit survival
#> probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Custom Time Points
By default, the function generates ~5 evenly spaced time points from
the conditioning time to the maximum follow-up. Use the
timePoints option to specify exact time points of clinical
interest (comma-separated string).
Clinically Meaningful Time Points
For example, 1-year, 2-year, 3-year, and 5-year conditional survival given survival to 12 months:
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 12,
timePoints = "24,36,48,60",
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultTime Points Including Values Before Conditioning Time
When a requested time point is at or before the conditioning time, the conditional survival is exactly 1.0 (the patient has already survived past that point).
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 24,
timePoints = "12,18,24,36,48,60",
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultSparse Time Points
Only two time points of interest:
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 6,
timePoints = "24,60",
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 24.0000000 6.0000000 0.5304933 ᵃ 0.07409589 0.38526807 0.6757186
#> Surgery 60.0000000 6.0000000 0.1805318 0.05796722 0.06691813 0.2941455
#> Surgery+Chemo 24.0000000 6.0000000 0.6909091 0.06772094 0.55817849 0.8236397
#> Surgery+Chemo 60.0000000 6.0000000 0.2841587 0.06070883 0.16517162 0.4031458
#> ─────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 6
#>
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 6.0 in the Surgery group (n at risk = 131), the estimated
#> conditional 60.0-unit survival probability was 18.1% (95% CI:
#> 6.7%--29.4%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 6.0 in the Surgery+Chemo group (n at
#> risk = 131), the estimated conditional 60.0-unit survival probability
#> was 28.4% (95% CI: 16.5%--40.3%).<p style='font-size:0.85em;
#> color:#666;'>Tip: Copy the text above directly into your manuscript
#> methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Factor Outcome Variable
The outcomeVar accepts both numeric (0/1) and factor
(2-level) variables. Factor variables are converted internally: the
first level becomes 0 (censored), the second level becomes 1
(event).
Using EventFactor (Alive/Dead)
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = EventFactor,
conditionVar = Treatment,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.34744752 0.69847084
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.10185033 0.38629006
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.00000000 0.08920005
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.45863838 0.79280941
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.23453082 0.53209882
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.07563188 0.34252974
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 0.00000000 0.26404973
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the Surgery group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 3.1% (95% CI:
#> 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Surgery+Chemo group (n
#> at risk = 110), the estimated conditional 117.8-unit survival
#> probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').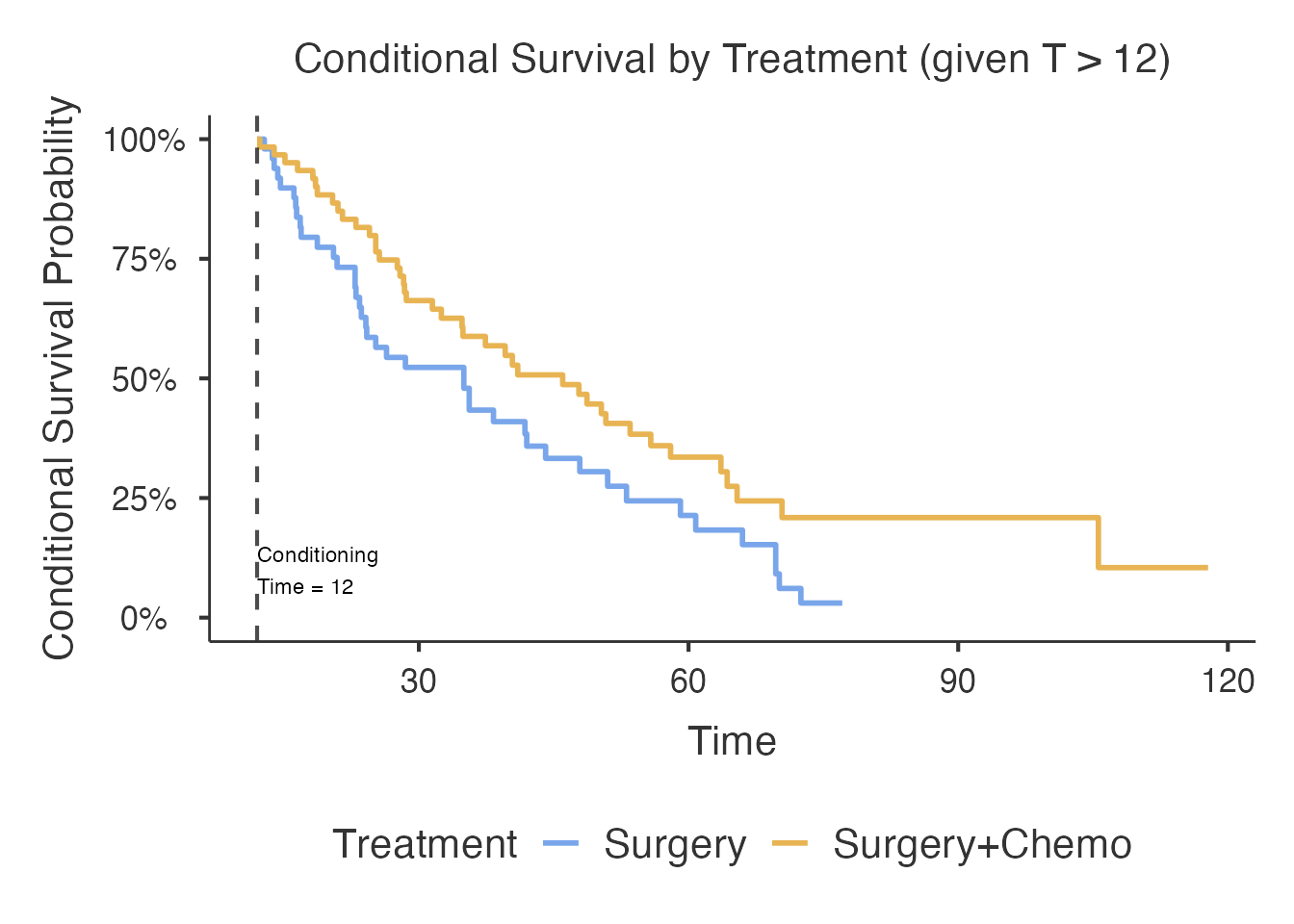
Confidence Level Variations
The confInt option controls the width of confidence
intervals around conditional survival estimates. The default is 0.95
(95% CI).
90% Confidence Intervals
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 12,
confInt = 0.90,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no default99% Confidence Intervals
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 12,
confInt = 0.99,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no default80% Confidence Intervals
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
confInt = 0.80,
method = "landmark",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Conditional Survival Probabilities
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Group Time Point Conditioning Time Conditional Survival Standard Error Lower CI Upper CI
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Surgery 33.2000000 12.0000000 0.52295918 ᵃ 0.08954841 0.4081983 0.63772009
#> Surgery 54.3000000 12.0000000 0.24407020 0.07256249 0.1510776 0.33706277
#> Surgery 75.5000000 12.0000000 0.03050877 0.02994508 0.0000000 0.06888494
#> Surgery 96.6000000 12.0000000 0.03050877 0.02994508 0.0000000 0.06888494
#> Surgery 117.8000000 12.0000000 0.03050877 0.02994508 0.0000000 0.06888494
#> Surgery+Chemo 33.2000000 12.0000000 0.62572389 0.08524928 0.5164725 0.73497524
#> Surgery+Chemo 54.3000000 12.0000000 0.38331482 0.07591160 0.2860302 0.48059945
#> Surgery+Chemo 75.5000000 12.0000000 0.20908081 0.06808744 0.1218232 0.29633838
#> Surgery+Chemo 96.6000000 12.0000000 0.20908081 0.06808744 0.1218232 0.29633838
#> Surgery+Chemo 117.8000000 12.0000000 0.10454041 0.08138380 2.428680e-4 0.20883795
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> ᵃ Conditional survival probabilities given survival to time 12
#>
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the landmark method. Given survival to
#> 12.0 in the Surgery group (n at risk = 110), the estimated conditional
#> 117.8-unit survival probability was 3.1% (80% CI: 0.0%--6.9%).
#>
#> Conditional survival analysis was performed using the landmark method.
#> Given survival to 12.0 in the Surgery+Chemo group (n at risk = 110),
#> the estimated conditional 117.8-unit survival probability was 10.5%
#> (80% CI: 0.0%--20.9%).<p style='font-size:0.85em; color:#666;'>Tip:
#> Copy the text above directly into your manuscript methods/results
#> section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Edge Cases
Conditioning Time at Median (Auto-Selection)
When conditionTime is 0, NULL, or NA, the function uses
the median follow-up time. This is the safest default for exploratory
analyses.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 0,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultLate Conditioning Time
A late conditioning time reduces the risk set. Estimates become less stable with wider confidence intervals.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 60,
method = "km",
showTable = TRUE,
showPlot = TRUE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultConditioning Time Beyond Maximum Follow-Up (Expect Error)
When the conditioning time is at or beyond the maximum observed follow-up, the function rejects the analysis with an informative error.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 200,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultToo Few Events (Expect Error)
A dataset with fewer than 5 events is rejected.
few_events_data <- data.frame(
time = c(5, 10, 15, 20, 25, 30, 35, 40, 45, 50,
55, 60, 65, 70, 75, 80, 85, 90, 95, 100),
event = c(1, 1, 0, 0, 0, 0, 0, 0, 0, 0,
0, 0, 0, 0, 0, 0, 0, 0, 0, 0),
group = rep(c("A", "B"), 10)
)
conditionalsurvival(
data = few_events_data,
timeVar = time,
outcomeVar = event,
conditionTime = 10,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultThree-Level Factor Outcome (Expect Error)
The outcome variable must have exactly 2 levels. A 3-level factor is rejected.
bad_data <- conditionalsurvival_test
bad_data$BadOutcome <- factor(
sample(c("Alive", "Dead", "Unknown"), nrow(bad_data), replace = TRUE)
)
conditionalsurvival(
data = bad_data,
timeVar = OverallTime,
outcomeVar = BadOutcome,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultSingle-Level Conditioning Variable (Expect Error)
A grouping variable with only one level cannot be used for stratified analysis.
single_data <- conditionalsurvival_test
single_data$SingleGroup <- "Everyone"
conditionalsurvival(
data = single_data,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = SingleGroup,
conditionTime = 12,
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error:
#> ! The conditioning variable has fewer than 2 levels. Stratified analysis requires at least 2 groups.Display Options: All Off
The showTable, showPlot, and
showExplanations toggles control visibility of their
respective panels. The todo, reportSentence,
and assumptions panels are always visible.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionVar = Treatment,
conditionTime = 12,
method = "km",
showTable = FALSE,
showPlot = FALSE,
showExplanations = FALSE
)
#>
#> CONDITIONAL SURVIVAL ESTIMATION
#>
#> Conditional Survival Estimation Analysis
#>
#> This analysis calculates conditional survival probabilities, which
#> represent the probability of surviving beyond a specific time point,
#> given survival to a conditioning time point.
#>
#> How to use this analysis:
#>
#> Time Variable: Select the survival time variable (numeric)Event/Status
#> Variable: Select the event indicator (0=censored, 1=event)Conditioning
#> Variable (optional): Variable for subgroup analysisSet Analysis
#> Options:Conditioning Time Point: Time at which to condition survival
#> (default: median follow-up)Estimation Method: Choose from Kaplan-Meier
#> weights, Landmark approach, IPW, or Presmoothed KMTime Points: Specify
#> comma-separated time points for analysis (e.g., 12,24,60)
#>
#> Interpretation:
#>
#> Conditional survival P(T > t | T > s) represents the probability of
#> surviving beyond time t, given survival to time s (conditioning time).
#> This is clinically relevant for patients who have already survived a
#> certain period and want to know their updated prognosis.
#>
#> Methods Available:
#>
#> Kaplan-Meier Weights: Uses weighted estimation with KM weightsLandmark
#> Approach: Subsets data to survivors at conditioning timeInverse
#> Probability Weighting: Accounts for censoring through
#> weightingPresmoothed KM: Smoothed version of Kaplan-Meier estimation
#>
#> Report Sentence (copy-ready)
#>
#> <div style='background-color:#f8f9fa; padding:10px; border-left:3px
#> solid #007bff; margin:8px 0; font-style:italic;'>Conditional survival
#> analysis was performed using the Kaplan-Meier weights method. Given
#> survival to 12.0 in the Surgery group (n at risk = 110), the estimated
#> conditional 117.8-unit survival probability was 3.1% (95% CI:
#> 0.0%--8.9%).
#>
#> Conditional survival analysis was performed using the Kaplan-Meier
#> weights method. Given survival to 12.0 in the Surgery+Chemo group (n
#> at risk = 110), the estimated conditional 117.8-unit survival
#> probability was 10.5% (95% CI: 0.0%--26.4%).<p
#> style='font-size:0.85em; color:#666;'>Tip: Copy the text above
#> directly into your manuscript methods/results section.
#>
#> Assumptions & Caveats
#>
#> Non-informative censoring: Censored patients must have the same future
#> survival probability as those who remain under
#> observation.Conditioning time: Must be within the observed follow-up
#> range. Estimates beyond maximum follow-up are unreliable.Sample size
#> at conditioning time: The number of patients still at risk at the
#> conditioning time must be adequate. Small risk sets produce unstable
#> estimates.Interpretation: CS(t|t0) = P(T > t | T > t0). This is NOT
#> the same as the unconditional survival probability.Clinical use:
#> Conditional survival is most useful for patient counseling after a
#> period of disease-free survival (e.g., '5-year survival given you
#> already survived 2 years').Plot Type Option (Currently Curves Only)
The plotType option accepts "curves",
"probability", or "both", but the plot
rendering currently always draws the curves style regardless of the
selected value.
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 12,
plotType = "probability",
method = "km",
showTable = FALSE,
showPlot = TRUE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultClinical Interpretation
Conditional survival is most valuable in clinical settings where patients return for follow-up visits and want updated prognostic information.
Key interpretation points:
CS(t | s) is always >= S(t): A patient who has survived to time s always has a survival probability at least as high as the original unconditional estimate at time t.
The conditioning effect is strongest early: The biggest prognostic “improvement” from conditional survival typically occurs in the first 1–3 years after treatment, when the hazard is highest.
Plateau behavior: If the hazard decreases over time (as in many cancers after successful treatment), conditional survival probabilities approach 1.0 as the conditioning time increases.
Group comparisons: Stratified conditional survival can reveal whether prognostic differences between treatment arms or stage groups narrow over time (a common pattern in oncology).
Clinical communication: The report sentence generated by this function is designed to be copied directly into manuscript results sections.
Example: Prognostic Update at Different Conditioning Times
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 6,
timePoints = "12,24,36,60",
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no default
conditionalsurvival(
data = conditionalsurvival_test,
timeVar = OverallTime,
outcomeVar = Event,
conditionTime = 24,
timePoints = "36,48,60,72",
method = "km",
showTable = TRUE,
showPlot = FALSE,
showExplanations = FALSE
)
#> Error in `conditionalsurvival()`:
#> ! argument "conditionVar" is missing, with no defaultReferences
Hieke S, Kleber M, Konig C, Engelhardt M, Schumacher M. Conditional survival: a useful concept to provide information on how prognosis evolves over time. Clin Cancer Res. 2015;21(7):1530-1536.
Zamboni BA, Yothers G, Choi M, et al. Conditional survival and the choice of conditioning set for patients with colon cancer: an analysis of NSABP trials C-03 through C-07. J Clin Oncol. 2010;28(15):2544-2548.
Skuladottir H, Olsen JH. Conditional survival of patients with the four main types of lung cancer in Denmark, 1978-2001. Cancer. 2003;97(8): 2014-2019.
Beran R. Nonparametric regression with randomly censored survival data. Technical Report, University of California, Berkeley. 1981.
Meira-Machado L, de Una-Alvarez J, Cadarso-Suarez C. Nonparametric estimation of transition probabilities in a non-Markov illness-death model. Lifetime Data Anal. 2006;12(3):325-344.