PLS Cox Regression for High-Dimensional Survival Analysis
Source:vignettes/jsurvival-plscox-comprehensive.Rmd
jsurvival-plscox-comprehensive.RmdNote: The
plscox()function is designed for use within jamovi’s GUI. The code examples below show the R syntax for reference. To run interactively, usedevtools::load_all()and call the R6 class directly:plscoxClass$new(options = plscoxOptions$new(...), data = mydata).
PLS Cox Regression
Overview
The PLS Cox Regression module (plscox)
combines Partial Least Squares (PLS) dimensionality reduction with Cox
proportional hazards modeling for high-dimensional survival data.
Instead of selecting individual variables like LASSO, PLS creates
latent components – weighted combinations of all
predictors that maximally explain covariance with the survival
outcome.
Key features:
- Component selection via cross-validation (log-likelihood, C-index), information criteria (BIC, AIC), or manual
- Multiple scaling methods (standardization, unit variance, min-max, none)
- Advanced PLS settings (sparse PLS, Q-squared stopping, p-value variable selection)
- Bootstrap validation (Harrell optimism-corrected C-index)
- Permutation testing for overall model significance
- Risk group stratification with Kaplan-Meier survival curves
- Variable importance (Cox-weighted PLS loadings)
- Data suitability assessment (traffic-light system with 6 checks)
PLS vs LASSO: Different Philosophies
| Approach | How It Works | Output |
|---|---|---|
| LASSO | Selects individual variables, drops others | Sparse coefficient vector |
| PLS | Creates weighted combinations of ALL variables | Component scores + loadings |
Use PLS when: - Variables are highly correlated (metabolomics, gene expression) - You want to retain information from all variables - The underlying signal comes from latent biological processes - p >> n and LASSO produces unstable selections
Use LASSO when: - You want to identify individual important predictors - A sparse, interpretable model is needed - Variables are relatively independent
How PLS Cox Works
- Extract PLS components: Find linear combinations of X that maximize covariance with the survival response
- Select components: Choose the optimal number via CV or information criteria
- Fit Cox model: Use selected components as predictors in Cox regression
- Interpret: Back-project component loadings to understand original variable contributions
Datasets Used in This Guide
| Dataset | N | Events | Predictors | Time Var | Status Var | Description |
|---|---|---|---|---|---|---|
plscox_metabolomics |
120 | ~50% | 80 metabolites + 3 clinical | survival_months |
death |
Metabolomic survival study with 3 latent pathways |
plscox_small |
50 | ~60% | 25 biomarkers | time_months |
status |
Small sample edge case |
plscox_genomic |
60 | ~55% | 200 genes | os_time |
os_event (numeric 0/1) |
True p>>n with missing data |
data_path2 <- "data/"
# Load metabolomics dataset (n=120, p=80)
load(paste0(data_path2, "plscox_metabolomics.rda"))
#> Error in `readChar()`:
#> ! cannot open the connection
cat("Metabolomics: N =", nrow(plscox_metabolomics),
"| Events =", sum(plscox_metabolomics$death == "Dead"),
"| Predictors =", sum(grepl("^METAB_", names(plscox_metabolomics))), "+ 3 clinical\n")
#> Metabolomics: N = 120 | Events = 69 | Predictors = 80 + 3 clinical
# Load small dataset (n=50, p=25)
load(paste0(data_path2, "plscox_small.rda"))
#> Error in `readChar()`:
#> ! cannot open the connection
cat("Small: N =", nrow(plscox_small),
"| Events =", sum(plscox_small$status == "Dead"),
"| Predictors =", sum(grepl("^MARKER_", names(plscox_small))), "\n")
#> Small: N = 50 | Events = 29 | Predictors = 25
# Load genomic dataset (n=60, p=200)
load(paste0(data_path2, "plscox_genomic.rda"))
#> Error in `readChar()`:
#> ! cannot open the connection
cat("Genomic: N =", nrow(plscox_genomic),
"| Events =", sum(plscox_genomic$os_event == 1),
"| Predictors =", sum(grepl("^GENE_", names(plscox_genomic))),
"| Missing values =", sum(is.na(plscox_genomic)), "\n")
#> Genomic: N = 60 | Events = 39 | Predictors = 200 | Missing values = 3581. Basic PLS Cox Analysis (Default Settings)
This demonstrates all default settings: 5 components, 10-fold CV, CV log-likelihood selection, standardization scaling, and all outputs/plots enabled.
metab_predictors <- c("age", "gender", "bmi",
paste0("METAB_", sprintf("%03d", 1:80)))
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
cross_validation = "k10",
component_selection = "cv_loglik",
scaling_method = "standardize",
suitabilityCheck = TRUE,
plot_components = TRUE,
plot_loadings = TRUE,
plot_scores = TRUE,
plot_validation = TRUE,
plot_survival = TRUE,
risk_groups = 3,
confidence_intervals = TRUE,
feature_importance = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetExpected: PLS should identify 2-3 important components reflecting the underlying pathway structure. Metabolites in blocks 1-15, 25-40, and 55-70 should have high loadings on the first few components.
2. Component Selection Methods
Cross-Validated Log-Likelihood (default)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
component_selection = "cv_loglik",
cross_validation = "k10",
plot_validation = TRUE,
feature_importance = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetCross-Validated C-Index
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
component_selection = "cv_cindex",
cross_validation = "k10",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetBIC (no cross-validation needed)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
component_selection = "bic",
cross_validation = "none",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetAIC
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
component_selection = "aic",
cross_validation = "none",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetManual (fixed number of components)
# Use exactly 3 components (based on domain knowledge)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 3,
component_selection = "manual",
cross_validation = "none",
plot_components = TRUE,
plot_loadings = TRUE,
feature_importance = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset3. Cross-Validation Methods
10-Fold CV (default)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
cross_validation = "k10",
component_selection = "cv_loglik",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset5-Fold CV (faster, slightly more bias)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
cross_validation = "k5",
component_selection = "cv_loglik",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetLeave-One-Out CV (small samples only)
small_predictors <- paste0("MARKER_", sprintf("%02d", 1:25))
plscox(
data = plscox_small,
time = "time_months",
status = "status",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = small_predictors,
pls_components = 5,
cross_validation = "loo",
component_selection = "cv_loglik",
plot_validation = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'small_predictors' which is not present in the dataset4. Scaling Methods
Variable scaling is critical for PLS since it operates on covariances.
Standardization (Z-scores, default)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
scaling_method = "standardize",
component_selection = "cv_loglik",
cross_validation = "k10",
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetUnit Variance Scaling (no centering)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
scaling_method = "unit_variance",
component_selection = "cv_loglik",
cross_validation = "k10",
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetMin-Max Scaling (range [0,1])
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
scaling_method = "minmax",
component_selection = "cv_loglik",
cross_validation = "k10",
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetNo Scaling
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
scaling_method = "none",
component_selection = "cv_loglik",
cross_validation = "k10",
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset5. Advanced PLS Settings
Sparse PLS (automatic variable selection within components)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
sparse_pls = TRUE,
component_selection = "cv_loglik",
cross_validation = "k10",
feature_importance = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetConvergence Tolerance
# Strict tolerance for higher precision
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
tolerance = 1e-10,
component_selection = "manual",
feature_importance = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetQ-Squared Limit (PLS stopping criterion)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
limQ2set = 0.5,
component_selection = "manual",
feature_importance = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetP-Value Based Variable Selection
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
pvals_expli = TRUE,
alpha_pvals_expli = 0.05,
component_selection = "manual",
feature_importance = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetTie Handling Method
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
tie_method = "breslow",
component_selection = "cv_loglik",
cross_validation = "k10",
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset6. Bootstrap Validation
Bootstrap validation assesses model overfitting using Harrell’s optimism-corrected C-index. Each bootstrap iteration: (1) fit model on bootstrap sample, (2) assess on bootstrap and original data, (3) compute optimism.
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
component_selection = "cv_loglik",
cross_validation = "k10",
bootstrap_validation = TRUE,
n_bootstrap = 100,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset7. Permutation Testing
Test whether the PLS components capture real survival signal or random patterns. The p-value is the proportion of permuted C-indices that exceed the original.
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
component_selection = "cv_loglik",
cross_validation = "k10",
permutation_test = TRUE,
n_permutations = 50, # Use 100+ for publication
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset8. Risk Stratification
Patients are stratified into risk groups based on quantiles of the PLS-derived linear predictor from the Cox model.
Binary Risk Groups
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 3,
component_selection = "manual",
risk_groups = 2,
plot_survival = TRUE,
confidence_intervals = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetQuartile Risk Groups
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 3,
component_selection = "manual",
risk_groups = 4,
plot_survival = TRUE,
confidence_intervals = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset9. Data Suitability Assessment
The traffic-light assessment checks 6 criteria: events-per-variable, reduction need, sample size, event rate, multicollinearity, and data quality.
# Metabolomics data: expected mostly green
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 3,
component_selection = "manual",
suitabilityCheck = TRUE,
plot_components = FALSE,
plot_loadings = FALSE,
plot_scores = FALSE,
plot_validation = FALSE,
plot_survival = FALSE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset
# Genomic p>>n data: expected yellow/red for EPV and data quality
gene_predictors <- paste0("GENE_", sprintf("%03d", 1:200))
plscox(
data = plscox_genomic,
time = "os_time",
status = "os_event",
outcomeLevel = "1",
censorLevel = "0",
predictors = gene_predictors,
pls_components = 5,
component_selection = "manual",
suitabilityCheck = TRUE,
plot_components = FALSE,
plot_loadings = FALSE,
plot_scores = FALSE,
plot_validation = FALSE,
plot_survival = FALSE
)
#> Error:
#> ! Argument 'predictors' contains 'gene_predictors' which is not present in the dataset10. Small Sample Analysis
plscox(
data = plscox_small,
time = "time_months",
status = "status",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = paste0("MARKER_", sprintf("%02d", 1:25)),
pls_components = 3,
cross_validation = "k5",
component_selection = "cv_loglik",
scaling_method = "standardize",
risk_groups = 2,
plot_components = TRUE,
plot_loadings = TRUE,
plot_survival = TRUE,
feature_importance = TRUE,
prediction_accuracy = TRUE,
suitabilityCheck = TRUE
)
#>
#> PARTIAL LEAST SQUARES COX MODELS
#>
#> <div style='background-color: #fff3cd; color: #856404; border: 1px
#> solid #ffeeba; padding: 12px; border-radius: 6px; margin-bottom:
#> 12px;'>Overall: Data is usable but review the flagged items.<table
#> style='width: 100%; border-collapse: collapse; font-size: 13px;'><tr
#> style='border-bottom: 2px solid #dee2e6;'><th style='padding: 6px;
#> text-align: left;'>Status<th style='padding: 6px; text-align:
#> left;'>Check<th style='padding: 6px; text-align: left;'>Value<th
#> style='padding: 6px; text-align: left;'>Detail<tr
#> style='border-bottom: 1px solid #dee2e6;'><td style='padding:
#> 6px;'><span style='color: #ffc107; font-size: 18px;'>●<td
#> style='padding: 6px;'>Events-Per-Variable (Overall)<td style='padding:
#> 6px;'>1.2 (n_events=29, p=25)<td style='padding: 6px;'>Low EPV for
#> standard modeling, but PLS handles this well through dimensionality
#> reduction.<tr style='border-bottom: 1px solid #dee2e6;'><td
#> style='padding: 6px;'><span style='color: #28a745; font-size:
#> 18px;'>●<td style='padding: 6px;'>Reduction Need<td
#> style='padding: 6px;'>p=25, n=50 (ratio=0.50)<td style='padding:
#> 6px;'>High-dimensional setting. Dimensionality reduction via PLS is
#> strongly indicated.<tr style='border-bottom: 1px solid #dee2e6;'><td
#> style='padding: 6px;'><span style='color: #ffc107; font-size:
#> 18px;'>●<td style='padding: 6px;'>Sample Size<td style='padding:
#> 6px;'>n=50<td style='padding: 6px;'>Small sample. Consider LOO
#> (Leave-One-Out) cross-validation instead of k-fold.<tr
#> style='border-bottom: 1px solid #dee2e6;'><td style='padding:
#> 6px;'><span style='color: #28a745; font-size: 18px;'>●<td
#> style='padding: 6px;'>Event Rate<td style='padding: 6px;'>58.0%
#> (29/50)<td style='padding: 6px;'>Balanced event rate. Good for model
#> estimation.<tr style='border-bottom: 1px solid #dee2e6;'><td
#> style='padding: 6px;'><span style='color: #28a745; font-size:
#> 18px;'>●<td style='padding: 6px;'>Multicollinearity<td
#> style='padding: 6px;'>Max |r| = 0.71<td style='padding: 6px;'>Moderate
#> collinearity. PLS effectively orthogonalizes these correlated
#> predictors.<tr style='border-bottom: 1px solid #dee2e6;'><td
#> style='padding: 6px;'><span style='color: #28a745; font-size:
#> 18px;'>●<td style='padding: 6px;'>Data Quality<td
#> style='padding: 6px;'>No missing data<td style='padding:
#> 6px;'>Complete dataset.
#>
#> PLS Cox Model Results
#>
#> Analysis Summary:
#>
#> Sample size: 50 subjects
#> Number of events: 29
#> Number of predictors: 25
#> PLS components used: 3
#> PLS algorithm: NIPALS (plsRcox default)
#> Component selection: Cross-Validated Log-Likelihood
#> Scaling method: standardize
#> Cross-validation: k5 (5 folds)
#> Tie handling: efron
#> Convergence tolerance: 1e-06
#> Q-squared limit: 0.0975
#>
#> Model Performance:
#>
#> Training Concordance Index: 0.783 (SE: 0.042)
#> Likelihood ratio test: 30.83 (p = 9.252e-07)
#> Wald test: 23.26 (p = 3.559e-05)
#>
#> Note: The Training Concordance Index overestimates true out-of-sample
#> performance, especially for high-dimensional data. Use Bootstrap or
#> Permutation tests for rigorous validation.
#>
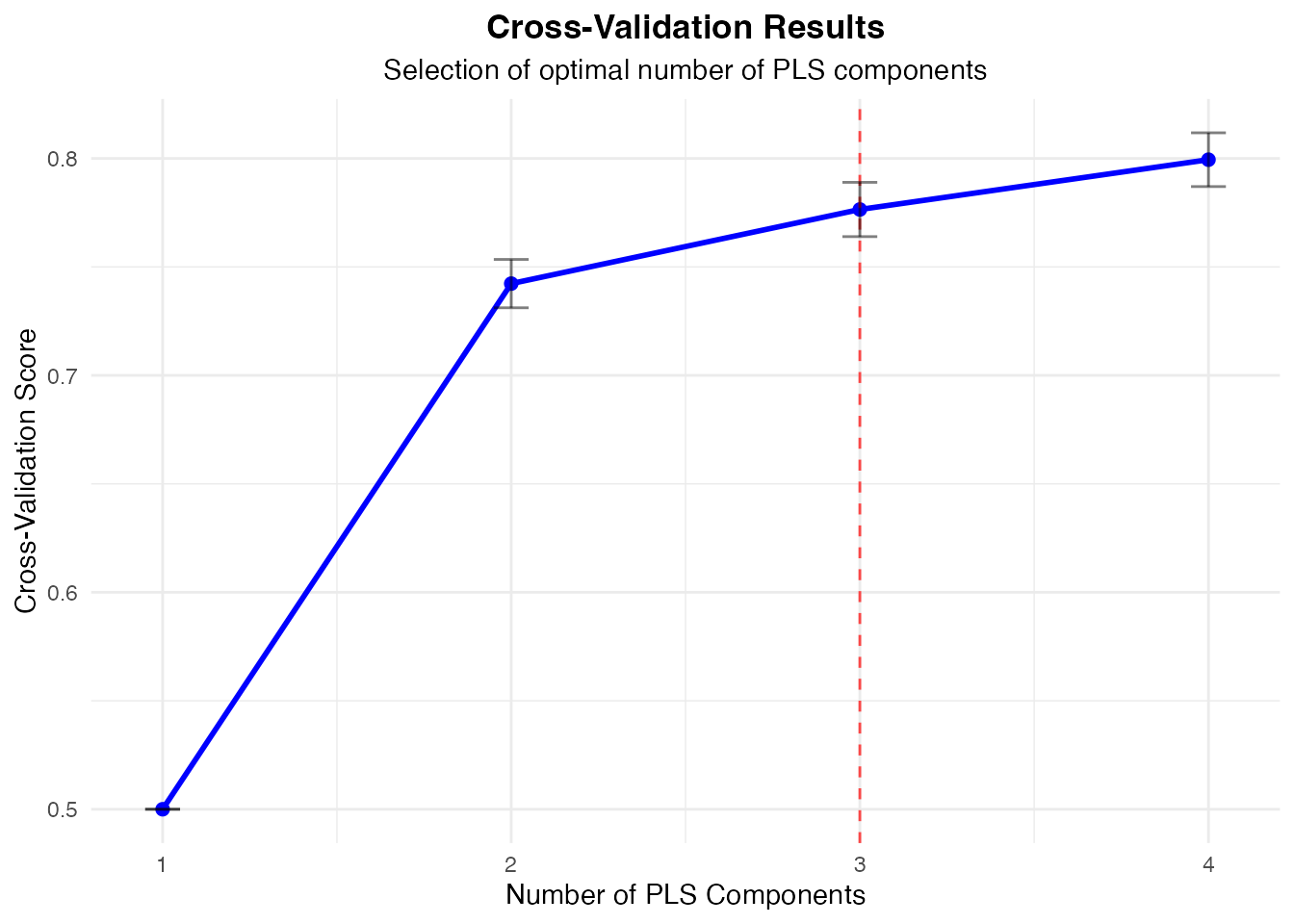
#> Component Selection Results
#> ──────────────────────────────────────────────────────────────────
#> Components CV Score SE C-Index Selected
#> ──────────────────────────────────────────────────────────────────
#> 1 0.5000000 1.570092e-17 No
#> 2 0.7423016 0.01114436 No
#> 3 0.7764710 0.01251187 Yes
#> 4 0.7994186 0.01239049 No
#> ──────────────────────────────────────────────────────────────────
#>
#>
#> PLS Cox Model Coefficients
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> PLS Component Coefficient Hazard Ratio HR Lower CI HR Upper CI Standard Error Z-value p-value
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> PLS Component 1 1.1897420 3.286233 2.0157554 5.357460 0.2493658 4.771071 0.0000018
#> PLS Component 2 0.4950034 1.640504 1.1480875 2.344118 0.1820982 2.718332 0.0065612
#> PLS Component 3 0.2705114 1.310635 0.9670626 1.776269 0.1551067 1.744035 0.0811530
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
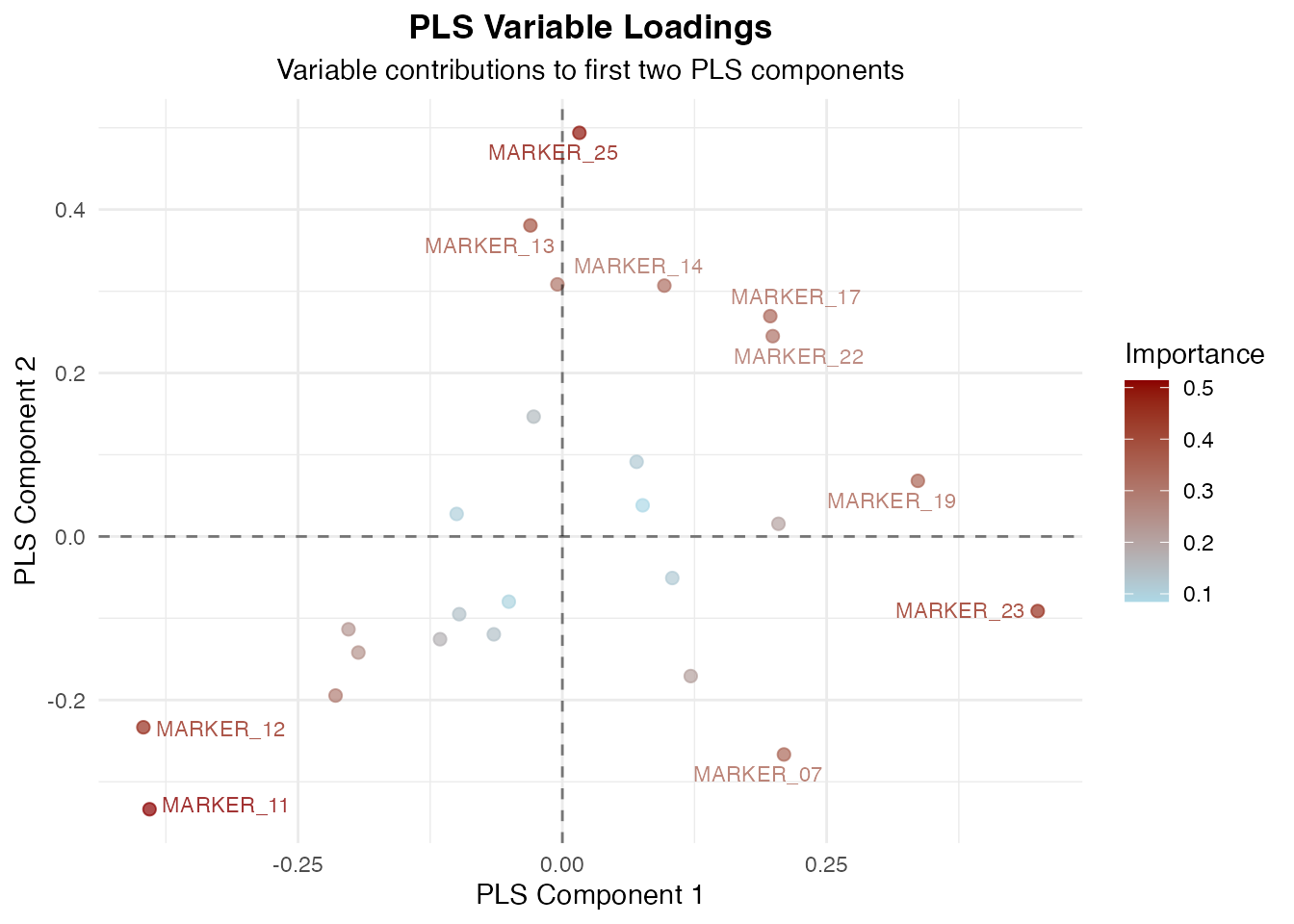
#> Variable Loadings on PLS Components
#> ───────────────────────────────────────────────────────────────────────────────────────
#> Variable Component 1 Component 2 Component 3 Cox-Weighted Importance
#> ───────────────────────────────────────────────────────────────────────────────────────
#> MARKER_23 0.449576752 -0.09121106 -0.365550228 0.6789156
#> MARKER_11 -0.390481147 -0.33365274 0.084881497 0.6526925
#> MARKER_12 -0.396282580 -0.23338378 -0.189486585 0.6382581
#> MARKER_19 0.336257540 0.06808261 0.162194848 0.4776364
#> MARKER_25 0.016111147 0.49379534 0.568336928 0.4173401
#> MARKER_07 0.209590451 -0.26665806 -0.115383327 0.4125677
#> MARKER_22 0.198997934 0.24505796 -0.165699557 0.4028843
#> MARKER_04 -0.214505528 -0.19456538 -0.078487980 0.3727487
#> MARKER_17 0.196649676 0.26954662 0.001737450 0.3678589
#> MARKER_14 0.096454668 0.30688689 0.343713734 0.3596447
#> MARKER_13 -0.030224409 0.38049243 0.430188221 0.3406751
#> MARKER_03 0.204375503 0.01555463 0.240185475 0.3158266
#> MARKER_15 -0.192965850 -0.14193953 -0.019314614 0.3050649
#> MARKER_18 -0.202310655 -0.11354504 0.005294904 0.2983350
#> MARKER_05 0.121415349 -0.17075152 0.239214527 0.2936858
#> MARKER_20 -0.097537150 -0.09504999 -0.413333938 0.2749056
#> MARKER_16 -0.115681010 -0.12567312 -0.277435819 0.2748887
#> MARKER_06 0.103966624 -0.05080671 0.186811961 0.1993777
#> MARKER_08 0.075949296 0.03812589 0.312833805 0.1938576
#> MARKER_10 -0.099993927 0.02758448 0.197968634 0.1861742
#> MARKER_21 -0.004697051 0.30841166 0.067719883 0.1765721
#> MARKER_02 -0.064839172 -0.11967818 0.082363952 0.1586634
#> MARKER_24 0.070225439 0.09135463 0.028021233 0.1363511
#> MARKER_09 -0.027147818 0.14660176 0.068839979 0.1234893
#> MARKER_01 -0.050641691 -0.07973672 0.017243703 0.1043851
#> ───────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Model Performance Metrics
#> ─────────────────────────────────────────────────────────────────────────────────────────
#> Metric Value Standard Error Lower CI Upper CI
#> ─────────────────────────────────────────────────────────────────────────────────────────
#> Training Concordance Index 0.7828283 0.04202365 0.7004619 0.8651946
#> R-squared (Nagelkerke) 0.4601716
#> AIC 179.0487826
#> BIC 184.7848516
#> ─────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Risk Group Stratification
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Risk Group N Subjects N Events Median Survival SE HR vs Low Risk p-value
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Risk Group 1 25 9 1.000000
#> Risk Group 2 25 20 6.800000 2.372449 3.860085 0.0009021
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Clinical Interpretation Guide
#>
#>
#>
#> PLS Components
#>
#> PLS components represent linear combinations of your original
#> predictors that are optimally related to survival outcomes. Each
#> component captures a different aspect of the biological variation in
#> your data.
#>
#>
#>
#> Variable Loadings
#>
#> Loadings indicate how much each original variable contributes to each
#> PLS component. Variables with higher absolute loadings have stronger
#> influence on that component.
#>
#>
#>
#> Hazard Ratios
#>
#> Note: The Hazard Ratios and p-values correspond to the abstract PLS
#> components, not your original variables. Each PLS component's hazard
#> ratio indicates the relative risk associated with a one-unit increase
#> in that component score. HR > 1 indicates increased risk, HR < 1
#> indicates decreased risk.
#>
#>
#>
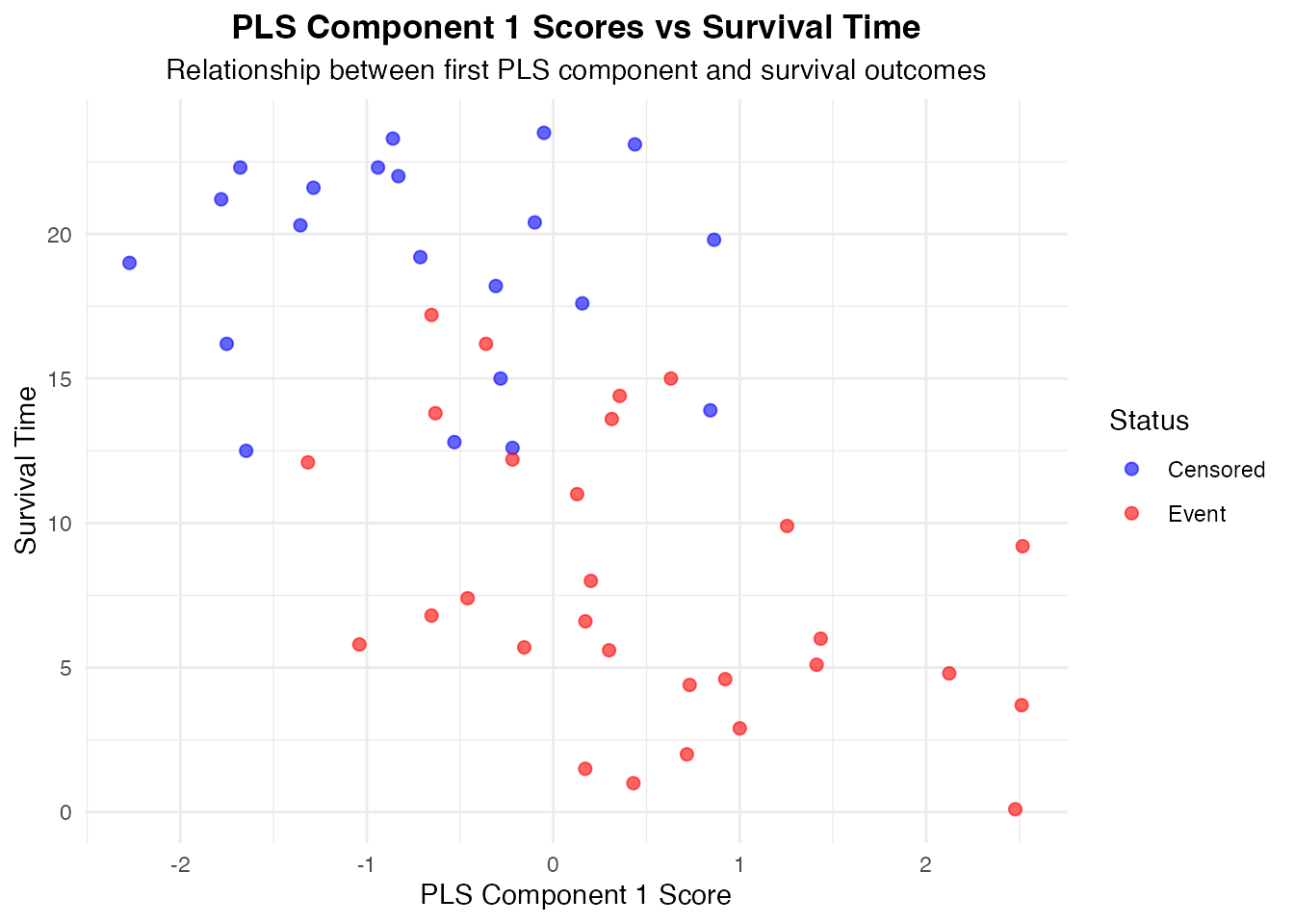
#> Risk Stratification
#>
#> Patients are grouped based on their overall PLS risk score. Higher
#> risk groups should show shorter survival times and more events.
#>
#>
#>
#> Model Validation
#>
#> Cross-validation helps select the optimal number of components.
#> Bootstrap validation can assess model stability and provide confidence
#> intervals for performance metrics.
#>
#>
#>
#> Clinical Application
#>
#> This model can be used for:
#>
#>
#> Risk stratification of patients
#> Identification of prognostic biomarker signatures
#> Treatment decision support
#> Clinical trial stratification
#>
#>
#>
#> Technical Notes and Assumptions
#>
#>
#>
#> PLS Cox Methodology
#>
#> This analysis combines Partial Least Squares (PLS) dimensionality
#> reduction with Cox proportional hazards regression. PLS finds
#> components that maximize covariance between predictors and the
#> survival outcome.
#>
#>
#>
#> Model Assumptions
#>
#>
#> Proportional Hazards: The hazard ratio for each component is constant
#> over time
#> Linear Relationships: Log-hazard is linear in the PLS components
#> Independence: Observations are independent
#> Non-informative Censoring: Censoring is independent of the event
#> process
#>
#>
#>
#>
#> Component Selection
#>
#> Cross-validation is used to select the optimal number of PLS
#> components to avoid overfitting while maintaining predictive
#> performance.
#>
#>
#>
#> Variable Scaling
#>
#> Predictor variables should be scaled when they have different units or
#> vastly different ranges to ensure fair contribution to PLS components.
#>
#>
#>
#> Sample Size Considerations
#>
#> For reliable results, aim for at least 10-15 events per PLS component
#> included in the model. With high-dimensional data, cross-validation
#> becomes crucial.
#>
#>
#>
#> Interpretation Cautions
#>
#>
#> PLS components are linear combinations - biological interpretation may
#> be complex
#> Variable importance should be interpreted in context of component
#> loadings
#> External validation is recommended before clinical application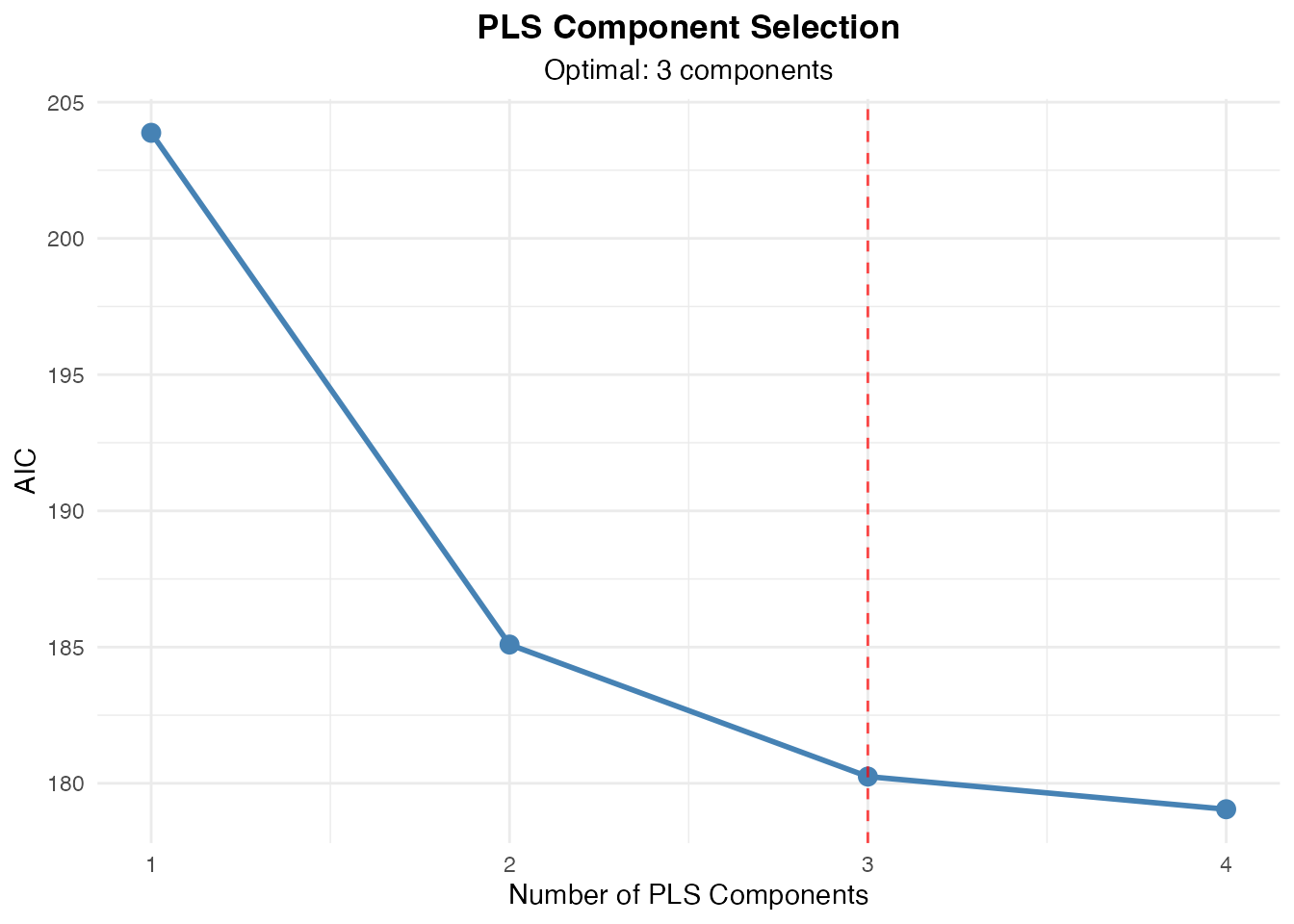
11. High-Dimensional Genomic Analysis (p >> n)
This is the core PLS use case: more genes than patients.
plscox(
data = plscox_genomic,
time = "os_time",
status = "os_event",
outcomeLevel = "1",
censorLevel = "0",
predictors = gene_predictors,
pls_components = 5,
component_selection = "bic",
cross_validation = "none",
scaling_method = "standardize",
risk_groups = 3,
plot_components = TRUE,
plot_loadings = TRUE,
plot_scores = TRUE,
plot_survival = TRUE,
feature_importance = TRUE,
prediction_accuracy = TRUE,
suitabilityCheck = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'gene_predictors' which is not present in the dataset12. All Plots Demonstration
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 5,
component_selection = "cv_loglik",
cross_validation = "k10",
risk_groups = 3,
plot_components = TRUE,
plot_loadings = TRUE,
plot_scores = TRUE,
plot_validation = TRUE,
plot_survival = TRUE,
feature_importance = TRUE,
confidence_intervals = TRUE,
prediction_accuracy = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset13. Full Validation Pipeline
Combine bootstrap validation and permutation testing for publication-quality results.
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 10,
component_selection = "cv_cindex",
cross_validation = "k5",
scaling_method = "standardize",
bootstrap_validation = TRUE,
n_bootstrap = 100,
permutation_test = TRUE,
n_permutations = 50,
risk_groups = 4,
plot_components = TRUE,
plot_loadings = TRUE,
plot_scores = TRUE,
plot_validation = TRUE,
plot_survival = TRUE,
confidence_intervals = TRUE,
feature_importance = TRUE,
prediction_accuracy = TRUE,
suitabilityCheck = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the dataset14. Edge Cases
Single Component Model
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 1,
component_selection = "manual",
plot_loadings = TRUE,
feature_importance = TRUE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetFew Predictors
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = c("age", "bmi"),
pls_components = 1,
component_selection = "manual",
feature_importance = TRUE,
prediction_accuracy = TRUE
)
#>
#> PARTIAL LEAST SQUARES COX MODELS
#>
#> <div style='background-color: #fff3cd; color: #856404; border: 1px
#> solid #ffeeba; padding: 12px; border-radius: 6px; margin-bottom:
#> 12px;'>Overall: Data is usable but review the flagged items.<table
#> style='width: 100%; border-collapse: collapse; font-size: 13px;'><tr
#> style='border-bottom: 2px solid #dee2e6;'><th style='padding: 6px;
#> text-align: left;'>Status<th style='padding: 6px; text-align:
#> left;'>Check<th style='padding: 6px; text-align: left;'>Value<th
#> style='padding: 6px; text-align: left;'>Detail<tr
#> style='border-bottom: 1px solid #dee2e6;'><td style='padding:
#> 6px;'><span style='color: #28a745; font-size: 18px;'>●<td
#> style='padding: 6px;'>Events-Per-Variable (Overall)<td style='padding:
#> 6px;'>34.5 (n_events=69, p=2)<td style='padding: 6px;'>High EPV. Model
#> estimation will be robust.<tr style='border-bottom: 1px solid
#> #dee2e6;'><td style='padding: 6px;'><span style='color: #ffc107;
#> font-size: 18px;'>●<td style='padding: 6px;'>Reduction Need<td
#> style='padding: 6px;'>p=2, EPV=34<td style='padding:
#> 6px;'>Moderate/low dimensionality. Standard Cox might suffice, but PLS
#> is still valid.<tr style='border-bottom: 1px solid #dee2e6;'><td
#> style='padding: 6px;'><span style='color: #28a745; font-size:
#> 18px;'>●<td style='padding: 6px;'>Sample Size<td style='padding:
#> 6px;'>n=120<td style='padding: 6px;'>Adequate sample size for PLS
#> regression cross-validation.<tr style='border-bottom: 1px solid
#> #dee2e6;'><td style='padding: 6px;'><span style='color: #28a745;
#> font-size: 18px;'>●<td style='padding: 6px;'>Event Rate<td
#> style='padding: 6px;'>57.5% (69/120)<td style='padding: 6px;'>Balanced
#> event rate. Good for model estimation.<tr style='border-bottom: 1px
#> solid #dee2e6;'><td style='padding: 6px;'><span style='color: #28a745;
#> font-size: 18px;'>●<td style='padding:
#> 6px;'>Multicollinearity<td style='padding: 6px;'>Max |r| = 0.04<td
#> style='padding: 6px;'>No concerning collinearity detected.<tr
#> style='border-bottom: 1px solid #dee2e6;'><td style='padding:
#> 6px;'><span style='color: #28a745; font-size: 18px;'>●<td
#> style='padding: 6px;'>Data Quality<td style='padding: 6px;'>No missing
#> data<td style='padding: 6px;'>Complete dataset.
#>
#> PLS Cox Model Results
#>
#> Analysis Summary:
#>
#> Sample size: 120 subjects
#> Number of events: 69
#> Number of predictors: 2
#> PLS components used: 1
#> PLS algorithm: NIPALS (plsRcox default)
#> Component selection: Manual
#> Scaling method: standardize
#> Cross-validation: k10 (10 folds)
#> Tie handling: efron
#> Convergence tolerance: 1e-06
#> Q-squared limit: 0.0975
#>
#> Model Performance:
#>
#> Training Concordance Index: 0.539 (SE: 0.04)
#> Likelihood ratio test: 1.34 (p = 0.2477)
#> Wald test: 1.33 (p = 0.2491)
#>
#> Note: The Training Concordance Index overestimates true out-of-sample
#> performance, especially for high-dimensional data. Use Bootstrap or
#> Permutation tests for rigorous validation.
#>
#> Component Selection Results
#> ───────────────────────────────────────────────────────
#> Components CV Score SE C-Index Selected
#> ───────────────────────────────────────────────────────
#> ───────────────────────────────────────────────────────
#> Note. Component selection: Manual. Using 1
#> component(s).
#>
#>
#> PLS Cox Model Coefficients
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> PLS Component Coefficient Hazard Ratio HR Lower CI HR Upper CI Standard Error Z-value p-value
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#> PLS Component 1 0.1417969 1.152343 0.9054498 1.466557 0.1230228 1.152606 0.2490721
#> ───────────────────────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
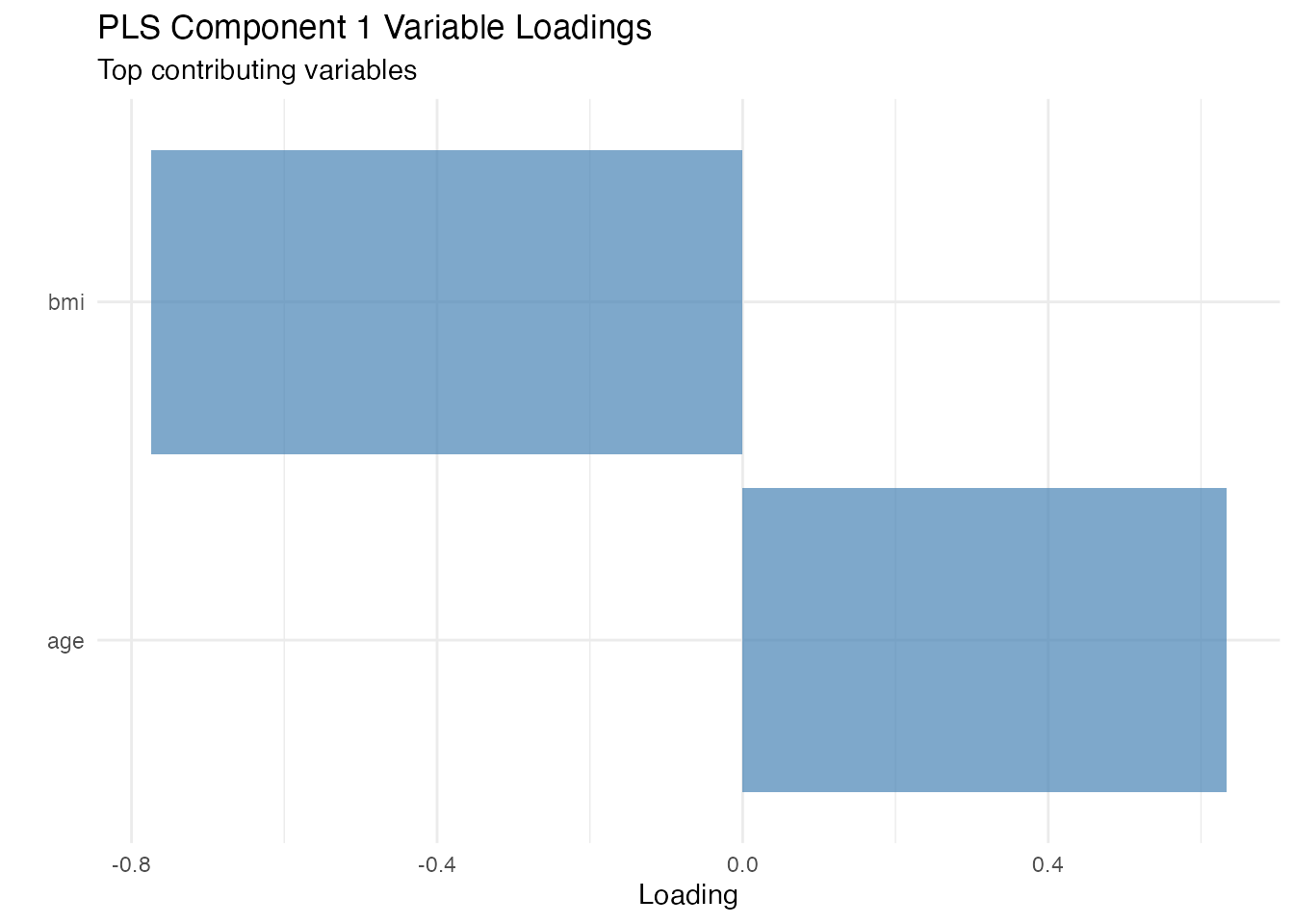
#> Variable Loadings on PLS Components
#> ────────────────────────────────────────────────────────────────────────────────────
#> Variable Component 1 Component 2 Component 3 Cox-Weighted Importance
#> ────────────────────────────────────────────────────────────────────────────────────
#> bmi -0.7742216 0.10978220
#> age 0.6329146 0.08974532
#> ────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Model Performance Metrics
#> ──────────────────────────────────────────────────────────────────────────────────────────
#> Metric Value Standard Error Lower CI Upper CI
#> ──────────────────────────────────────────────────────────────────────────────────────────
#> Training Concordance Index 0.53922040 0.04034305 0.4601480 0.6182928
#> R-squared (Nagelkerke) 0.01107350
#> AIC 542.39315135
#> BIC 545.18064309
#> ──────────────────────────────────────────────────────────────────────────────────────────
#>
#>
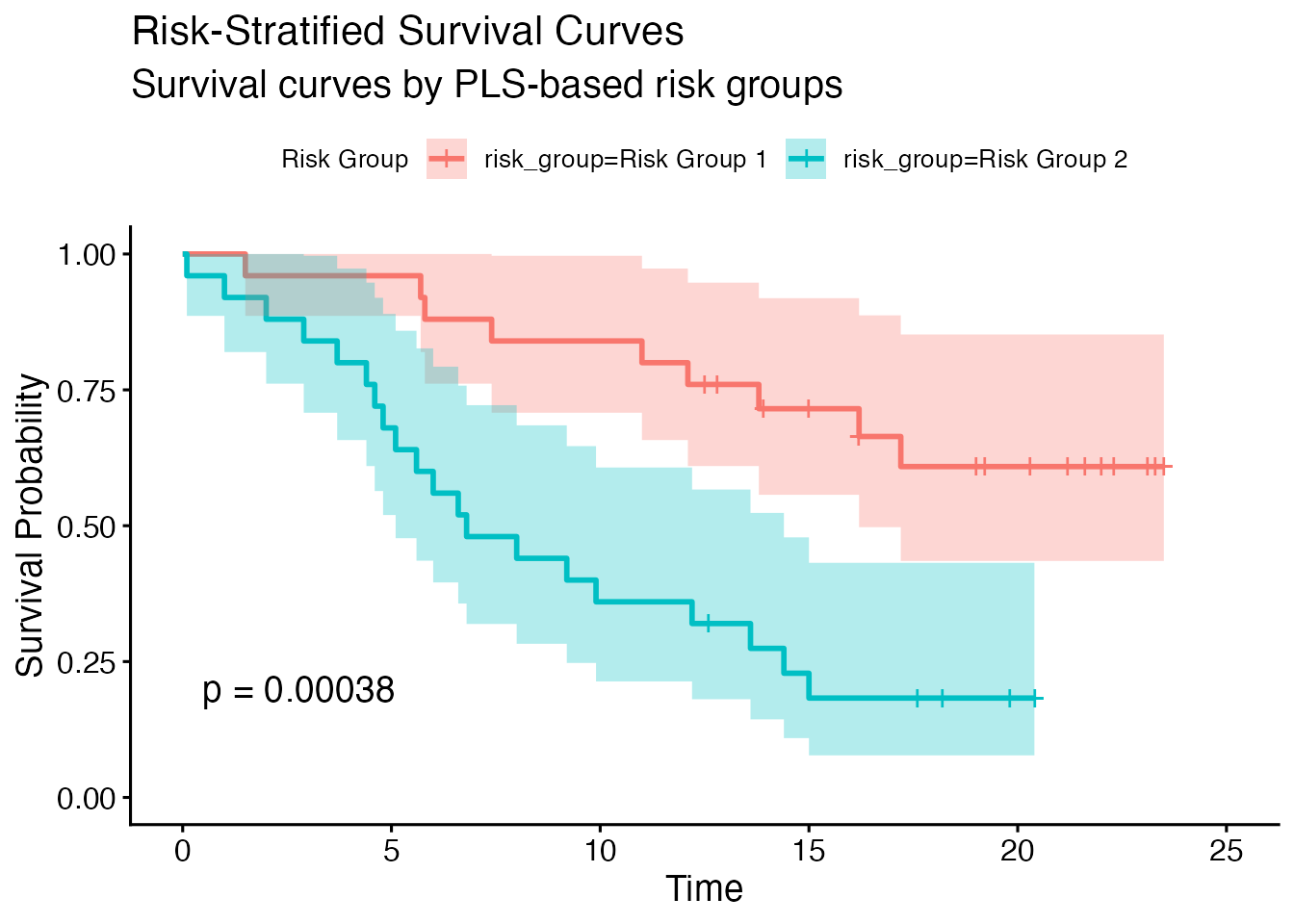
#> Risk Group Stratification
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Risk Group N Subjects N Events Median Survival SE HR vs Low Risk p-value
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#> Risk Group 1 40 19 16.90000 1.000000
#> Risk Group 2 40 26 16.00000 3.265306 1.445136 0.2281569
#> Risk Group 3 40 24 19.10000 4.540816 1.350071 0.3299992
#> ────────────────────────────────────────────────────────────────────────────────────────────────────────
#>
#>
#> Clinical Interpretation Guide
#>
#>
#>
#> PLS Components
#>
#> PLS components represent linear combinations of your original
#> predictors that are optimally related to survival outcomes. Each
#> component captures a different aspect of the biological variation in
#> your data.
#>
#>
#>
#> Variable Loadings
#>
#> Loadings indicate how much each original variable contributes to each
#> PLS component. Variables with higher absolute loadings have stronger
#> influence on that component.
#>
#>
#>
#> Hazard Ratios
#>
#> Note: The Hazard Ratios and p-values correspond to the abstract PLS
#> components, not your original variables. Each PLS component's hazard
#> ratio indicates the relative risk associated with a one-unit increase
#> in that component score. HR > 1 indicates increased risk, HR < 1
#> indicates decreased risk.
#>
#>
#>
#> Risk Stratification
#>
#> Patients are grouped based on their overall PLS risk score. Higher
#> risk groups should show shorter survival times and more events.
#>
#>
#>
#> Model Validation
#>
#> Cross-validation helps select the optimal number of components.
#> Bootstrap validation can assess model stability and provide confidence
#> intervals for performance metrics.
#>
#>
#>
#> Clinical Application
#>
#> This model can be used for:
#>
#>
#> Risk stratification of patients
#> Identification of prognostic biomarker signatures
#> Treatment decision support
#> Clinical trial stratification
#>
#>
#>
#> Technical Notes and Assumptions
#>
#>
#>
#> PLS Cox Methodology
#>
#> This analysis combines Partial Least Squares (PLS) dimensionality
#> reduction with Cox proportional hazards regression. PLS finds
#> components that maximize covariance between predictors and the
#> survival outcome.
#>
#>
#>
#> Model Assumptions
#>
#>
#> Proportional Hazards: The hazard ratio for each component is constant
#> over time
#> Linear Relationships: Log-hazard is linear in the PLS components
#> Independence: Observations are independent
#> Non-informative Censoring: Censoring is independent of the event
#> process
#>
#>
#>
#>
#> Component Selection
#>
#> Cross-validation is used to select the optimal number of PLS
#> components to avoid overfitting while maintaining predictive
#> performance.
#>
#>
#>
#> Variable Scaling
#>
#> Predictor variables should be scaled when they have different units or
#> vastly different ranges to ensure fair contribution to PLS components.
#>
#>
#>
#> Sample Size Considerations
#>
#> For reliable results, aim for at least 10-15 events per PLS component
#> included in the model. With high-dimensional data, cross-validation
#> becomes crucial.
#>
#>
#>
#> Interpretation Cautions
#>
#>
#> PLS components are linear combinations - biological interpretation may
#> be complex
#> Variable importance should be interpreted in context of component
#> loadings
#> External validation is recommended before clinical application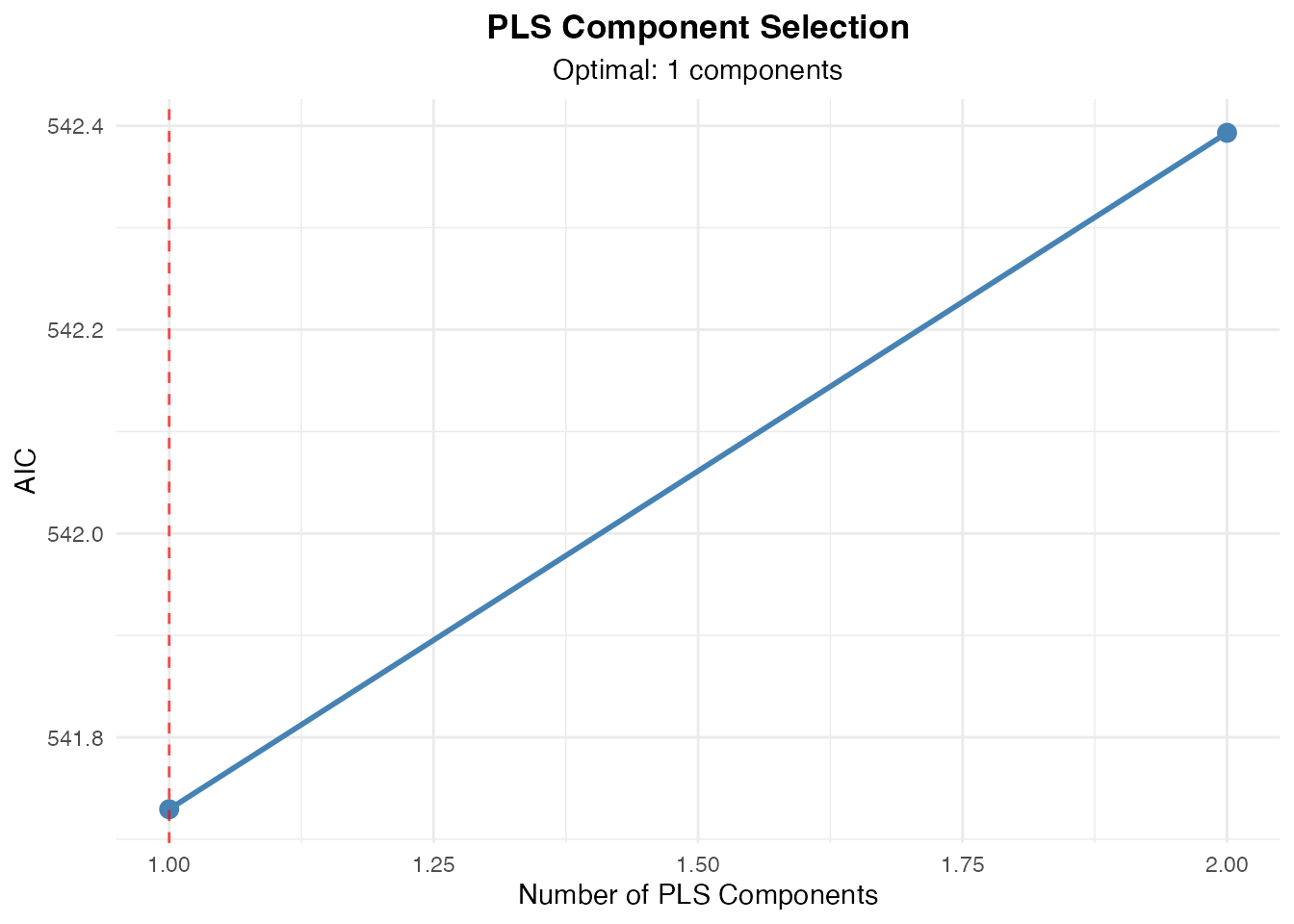
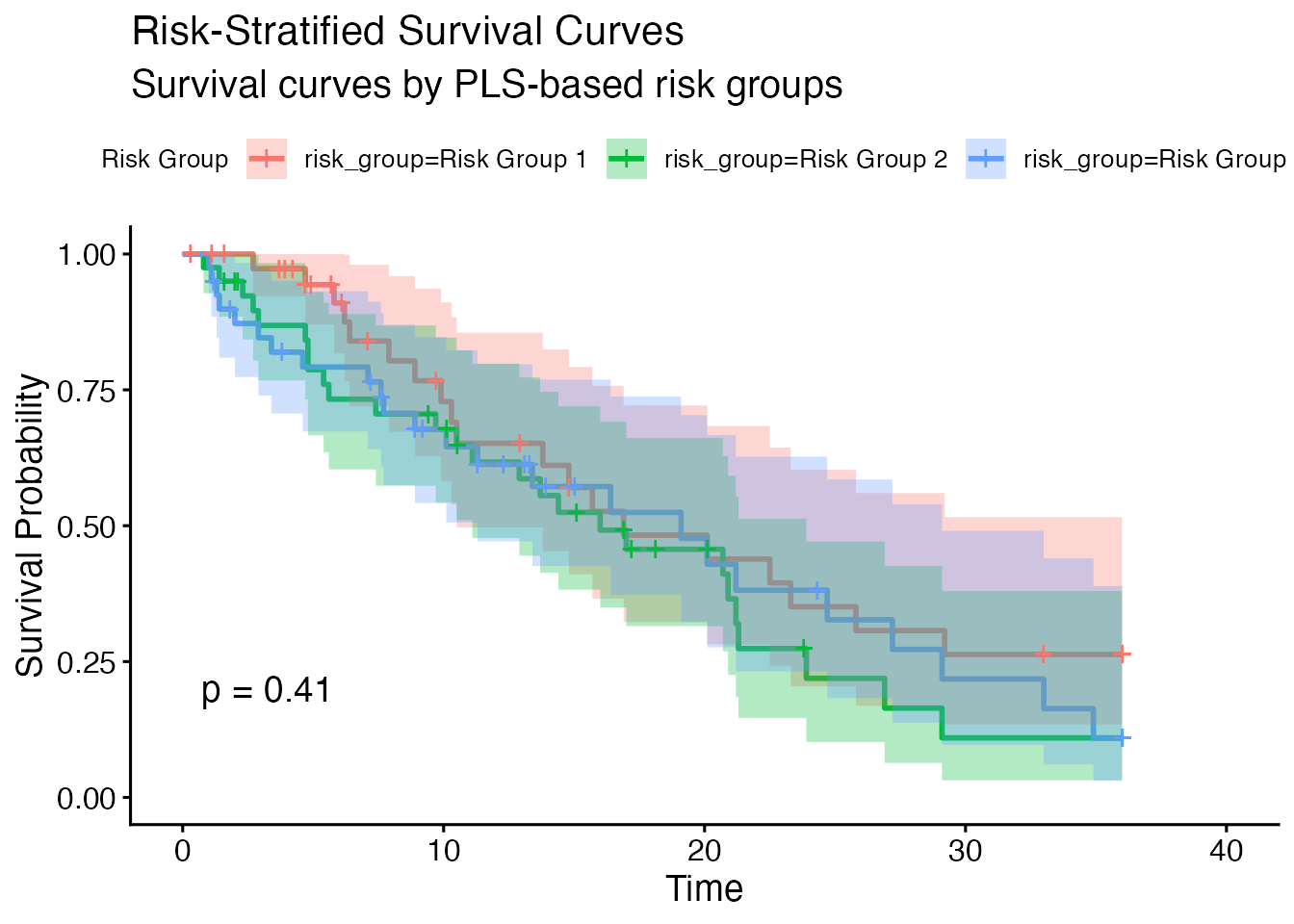
Minimal Output (no plots, no optional tables)
plscox(
data = plscox_metabolomics,
time = "survival_months",
status = "death",
outcomeLevel = "Dead",
censorLevel = "Alive",
predictors = metab_predictors,
pls_components = 3,
component_selection = "manual",
suitabilityCheck = FALSE,
plot_components = FALSE,
plot_loadings = FALSE,
plot_scores = FALSE,
plot_validation = FALSE,
plot_survival = FALSE,
confidence_intervals = FALSE,
feature_importance = FALSE,
prediction_accuracy = FALSE,
bootstrap_validation = FALSE,
permutation_test = FALSE
)
#> Error:
#> ! Argument 'predictors' contains 'metab_predictors' which is not present in the datasetInterpreting Results
Model Summary
The Model Summary HTML shows: sample size, events, predictors, number of PLS components used, selection method, scaling, CV method, tie handling, tolerance, and training C-index with likelihood ratio and Wald tests.
Component Selection Table
| Column | Meaning |
|---|---|
| Components | Number of PLS components considered |
| CV Score | Cross-validation score (log-likelihood, C-index, AIC, or BIC) |
| SE | Standard error of CV score (when available) |
| C-Index | Concordance index for that number of components (AIC/BIC only) |
| Selected | “Yes” marks the optimal number of components |
Model Coefficients Table
| Column | Meaning |
|---|---|
| PLS Component | Component identifier (PLS_1, PLS_2, …) |
| Coefficient | Cox regression coefficient |
| Hazard Ratio | exp(coefficient) |
| HR Lower CI / HR Upper CI | 95% confidence interval for HR |
| Standard Error | SE of the coefficient |
| Z-value | Wald test statistic |
| p-value | Significance of component in Cox model |
PLS vs PCA for Survival Analysis
| Method | Supervision | Components Maximize |
|---|---|---|
| PCA | Unsupervised | Variance in X only |
| PLS | Supervised | Covariance between X and Y (survival) |
PLS components are specifically constructed to predict survival, while PCA components may capture variance unrelated to the outcome.
Common Pitfalls
Too many components: More components can mean overfitting. Always use cross-validation to select the optimal number.
Ignoring variable scaling: Metabolites on different scales dominate PLS if not standardized. Always use
scaling_method = "standardize"(the default).Not validating: Use bootstrap validation and/or permutation testing to assess reliability. The training C-index overestimates true performance.
Interpreting loadings as independent effects: A high loading means the variable contributes to a component, not that it has an independent causal effect. Groups of correlated variables share loading magnitude.
Using LOO CV for large datasets: Leave-one-out is computationally expensive and can be unstable. Use 5- or 10-fold CV for n > 50.
Sparse PLS with too few components: Sparse PLS may return NULL component scores for some configurations. If this happens, try increasing components or disabling sparse mode.
Related ClinicoPath Functions
| Function | Use When |
|---|---|
LASSO Cox (lassocox) |
Want sparse individual variable selection |
Adaptive LASSO (adaptivelasso) |
Oracle property variable selection |
NCV Reg Cox (ncvregcox) |
SCAD/MCP non-convex penalties |
High-Dimensional Cox (highdimcox) |
Multiple regularization methods |
PCA Cox (pcacox) |
Unsupervised dimensionality reduction |
| Multivariable Survival | Standard Cox with few predictors |
References
- Bastien P, Esposito Vinzi V, Tenenhaus M. PLS generalised linear regression. Comput Stat Data Anal. 2005;48(1):17-46.
- Boulesteix AL, Strimmer K. Partial least squares: a versatile tool for the analysis of high-dimensional genomic data. Brief Bioinform. 2007;8(1):32-44.
- Li H, Gui J. Partial Cox regression analysis for high-dimensional microarray gene expression data. Bioinformatics. 2004;20(Suppl 1):i208-i215.
- Mevik BH, Wehrens R. The pls package: principal component and partial least squares regression in R. J Stat Softw. 2007;18(2):1-23.